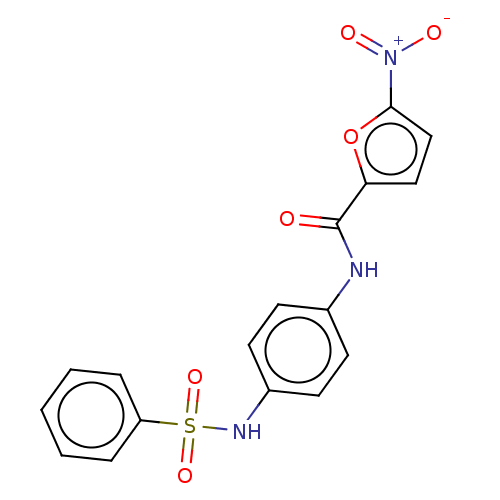
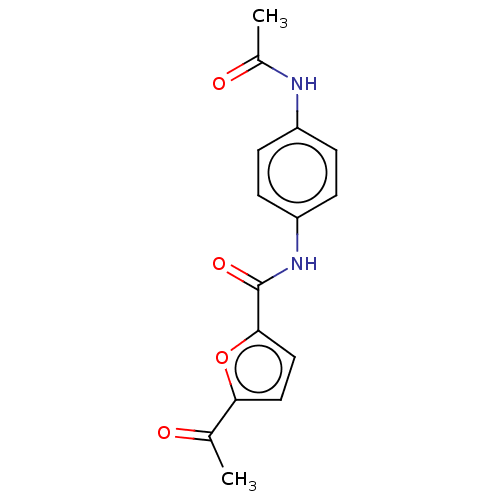
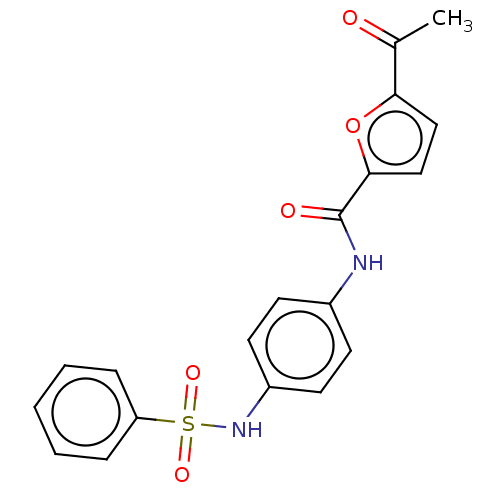
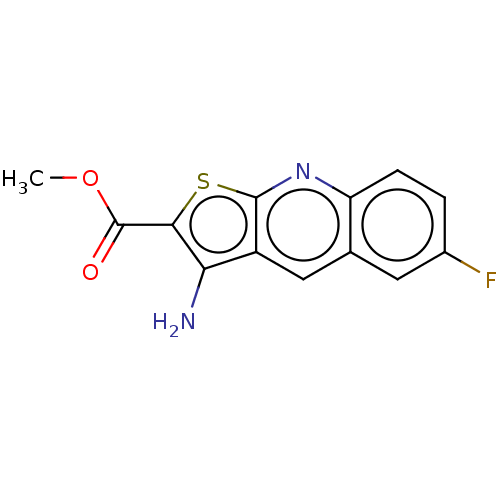
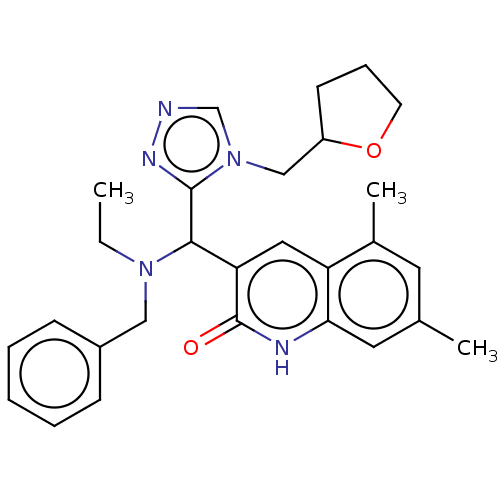
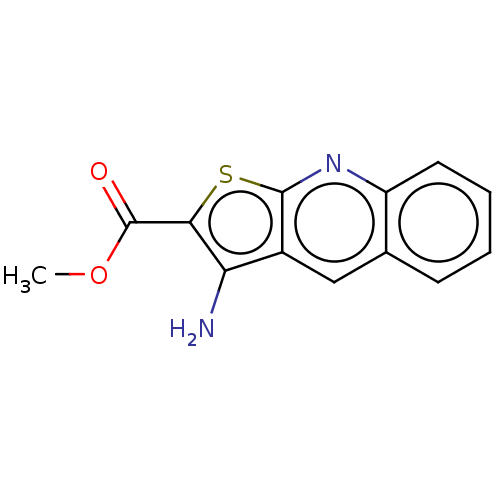
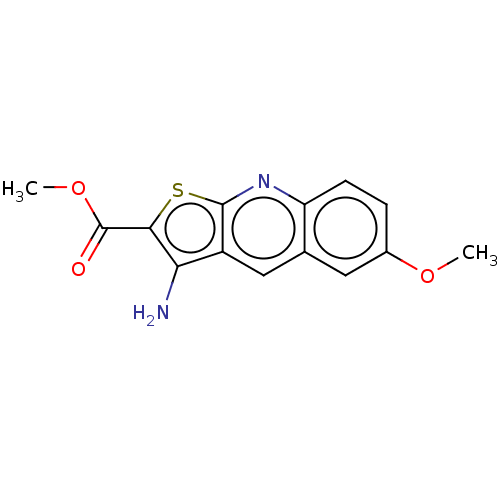
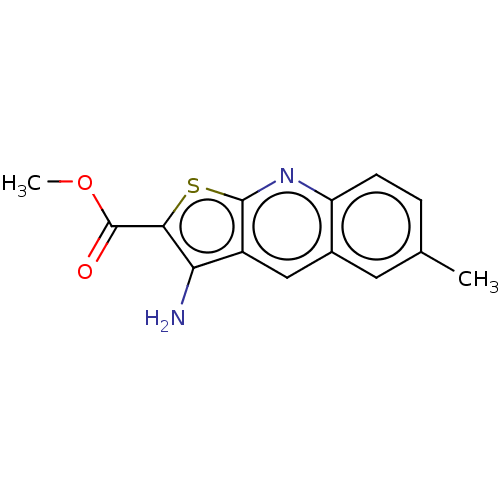
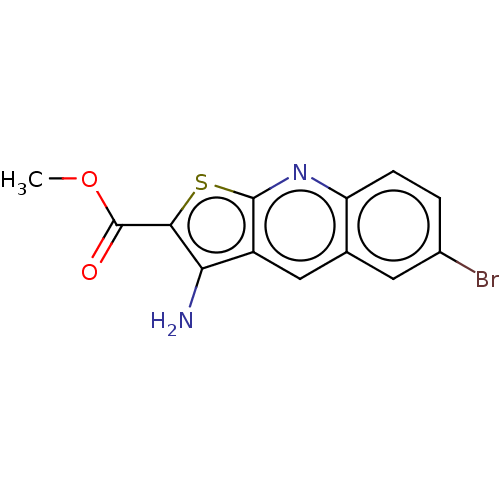
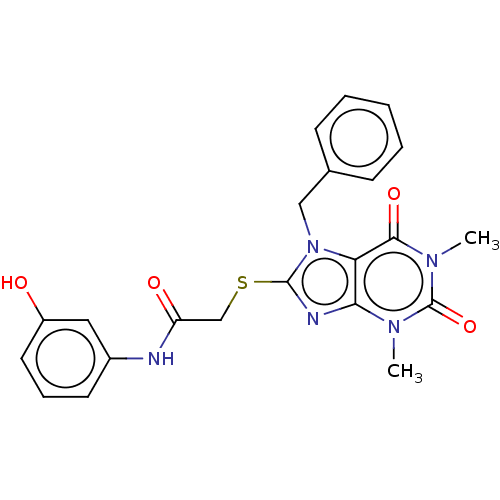
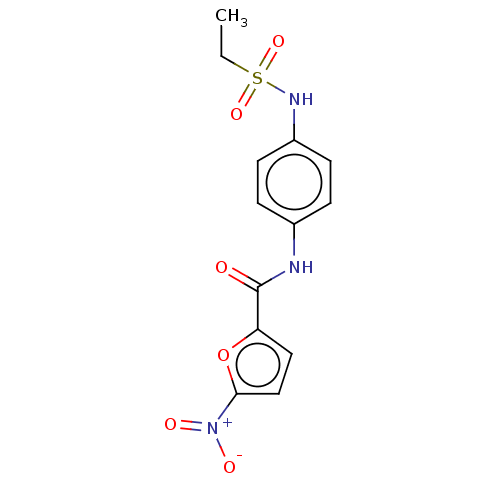
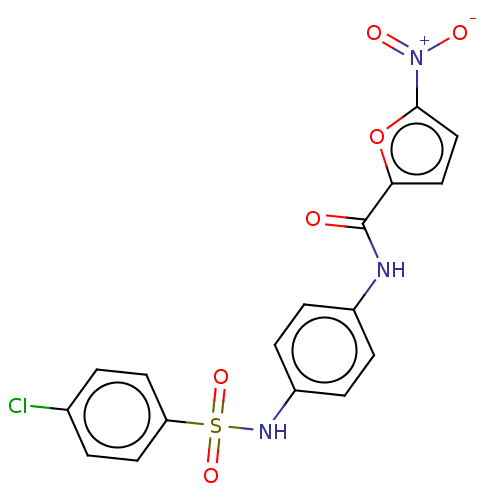
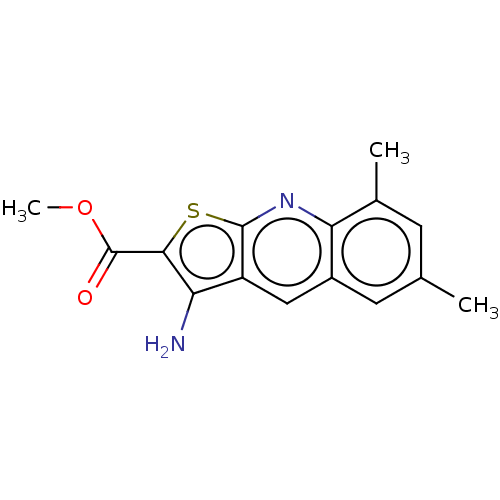
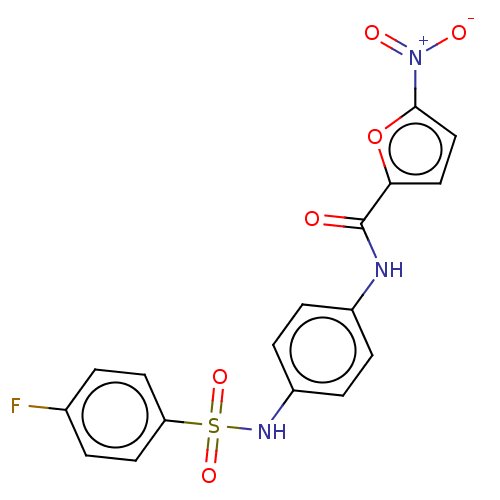
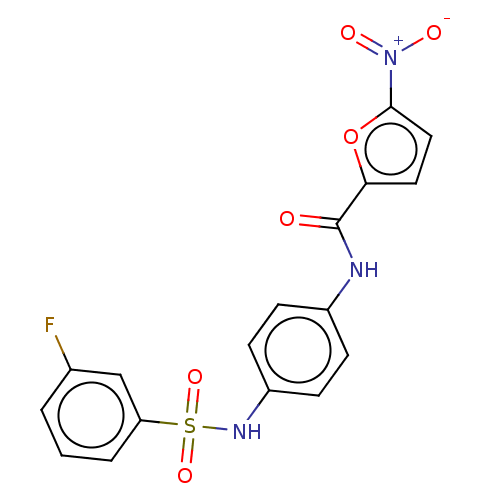
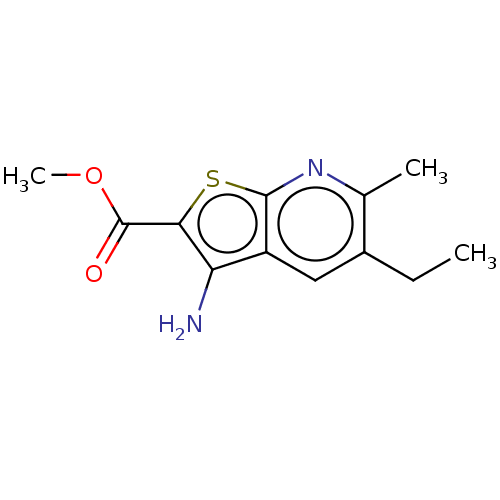
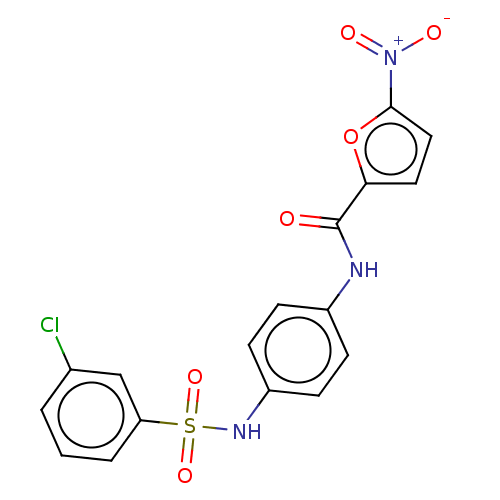
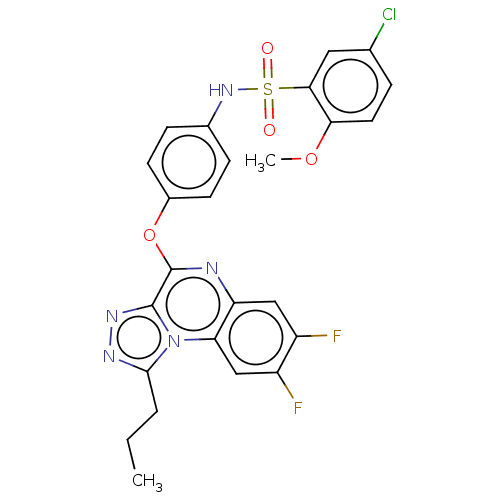
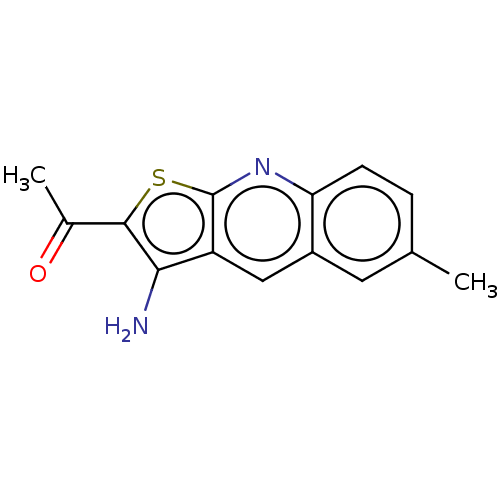
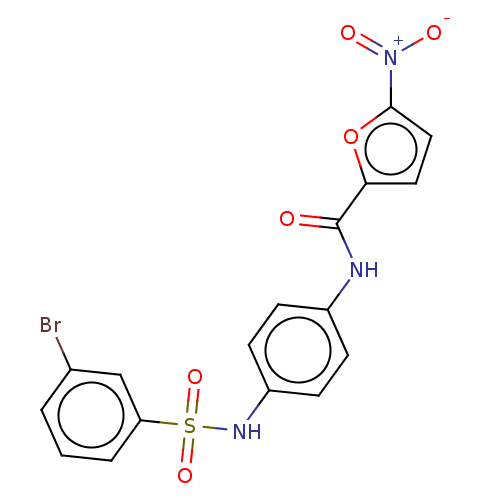
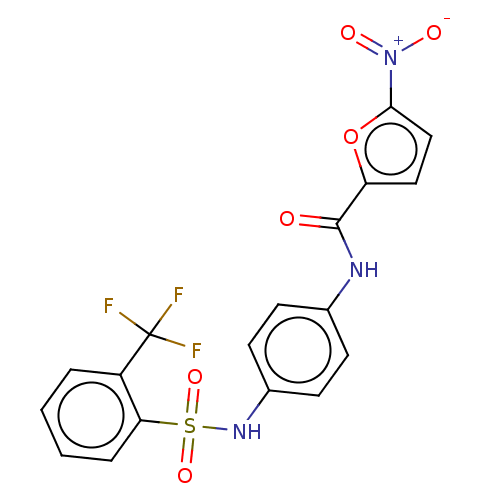
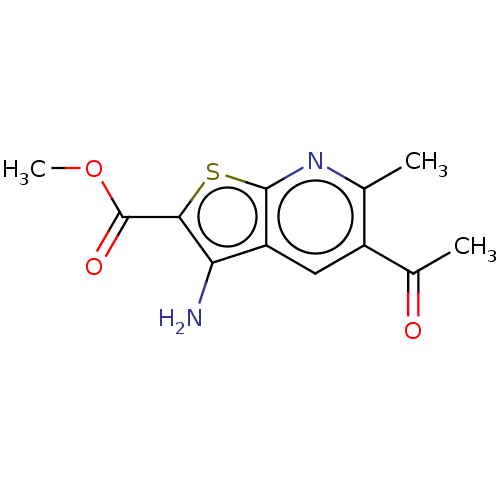
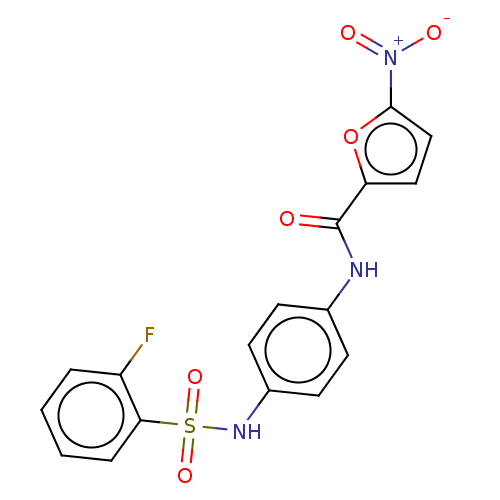
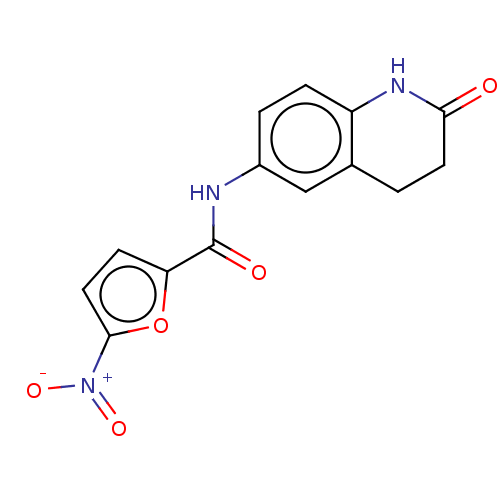
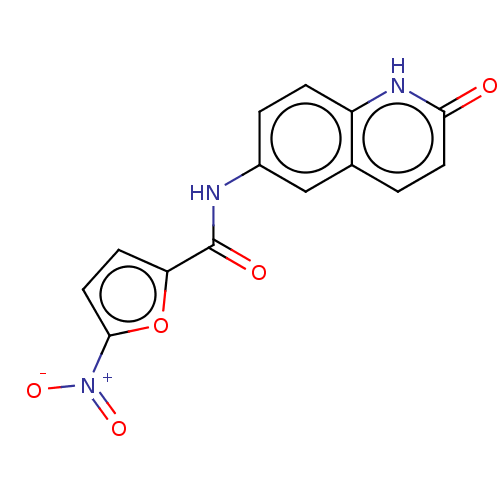
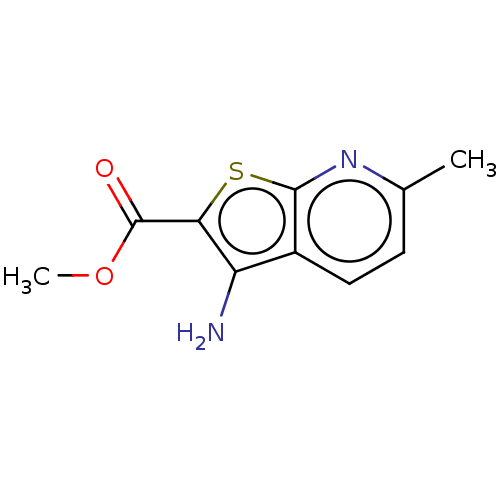
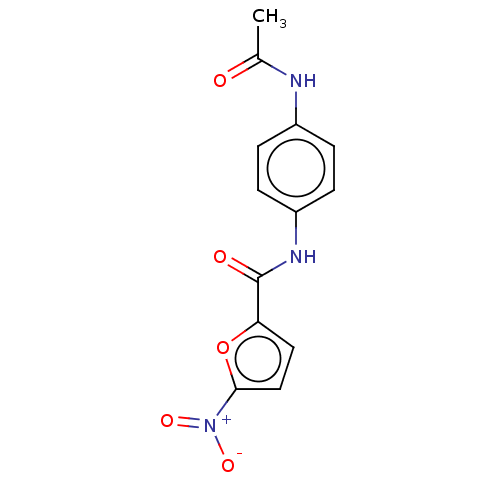
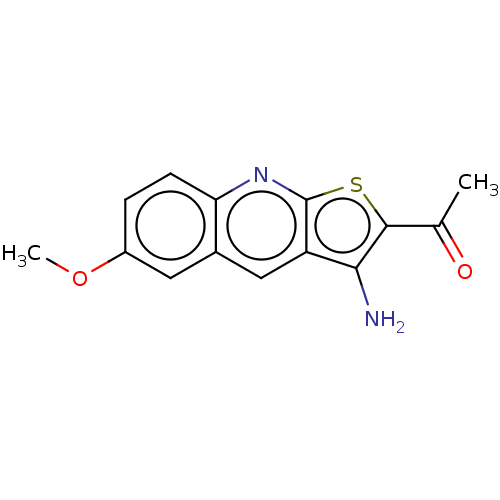
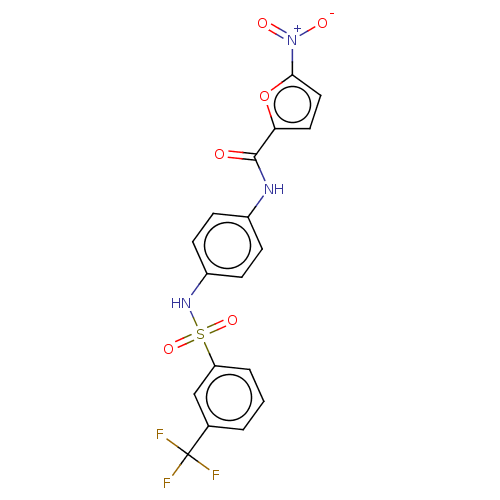
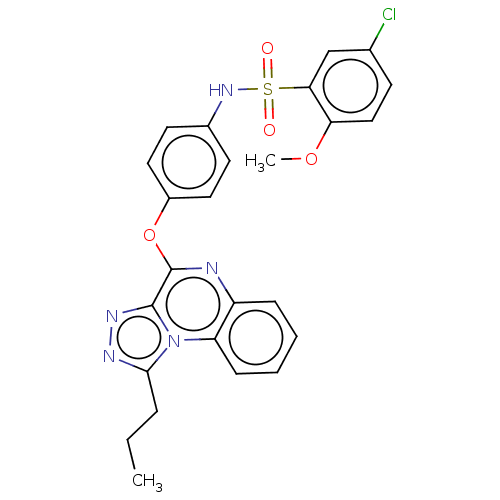
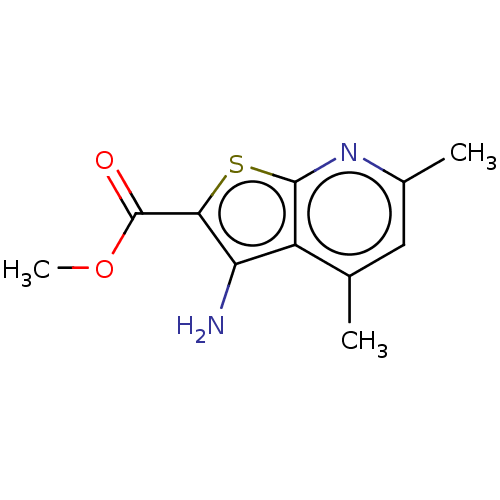
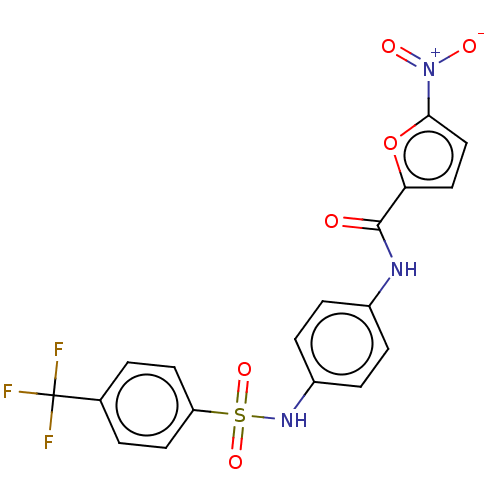
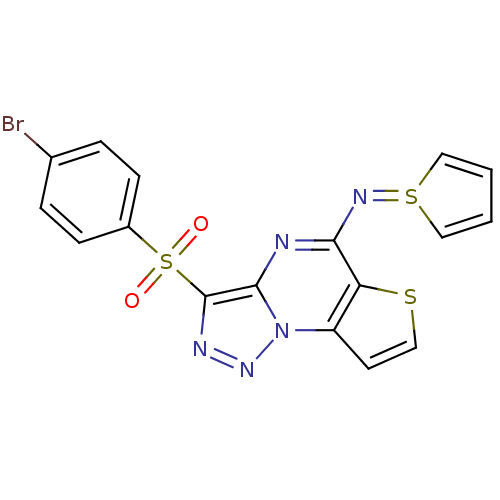
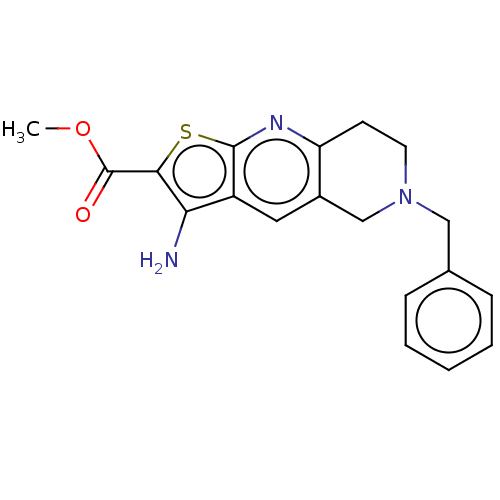
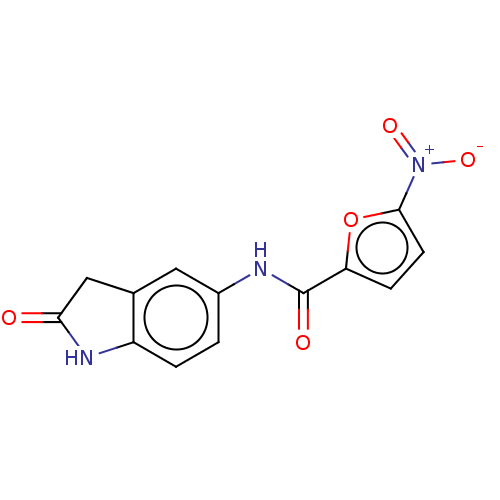
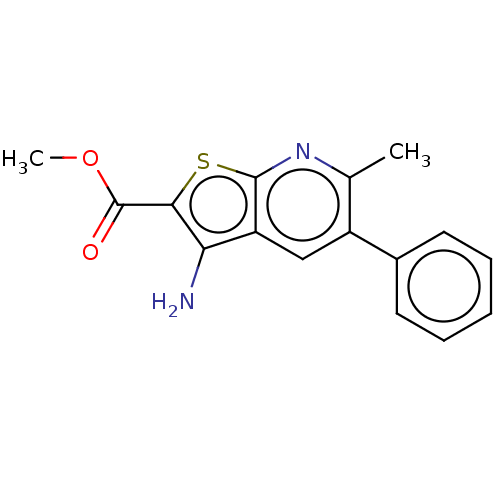
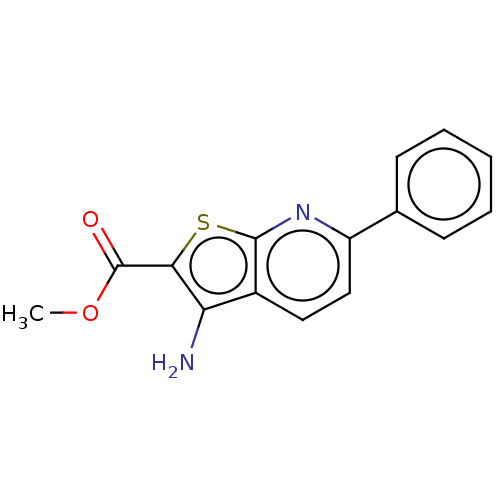
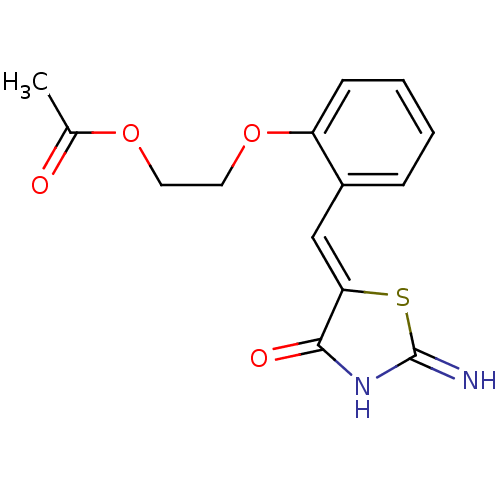
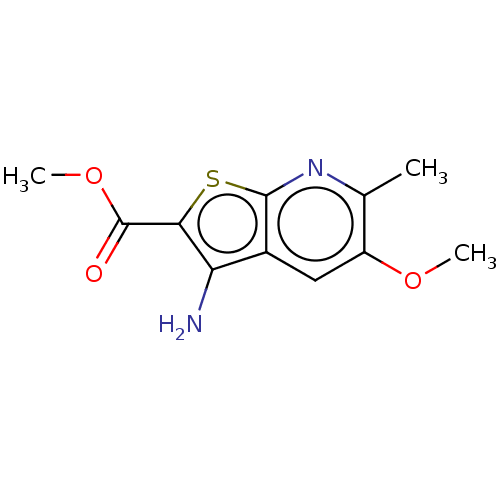
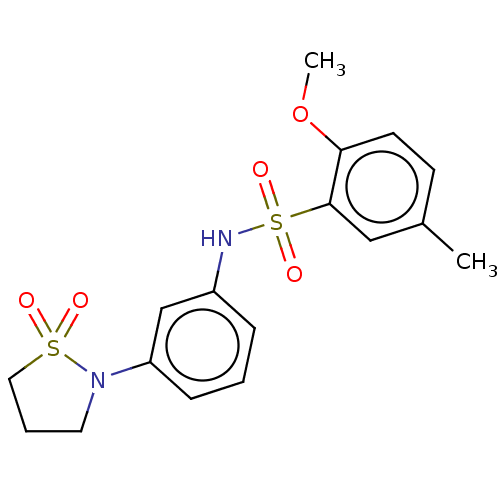
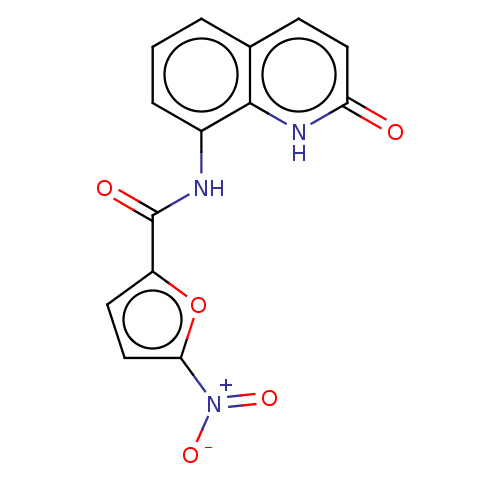
Affinity DataIC50: 110nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
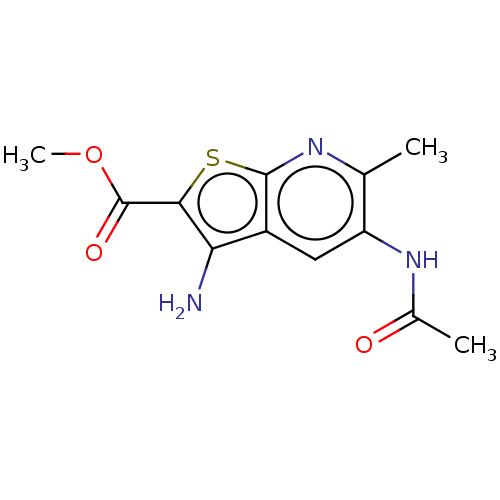
Affinity DataIC50: 140nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
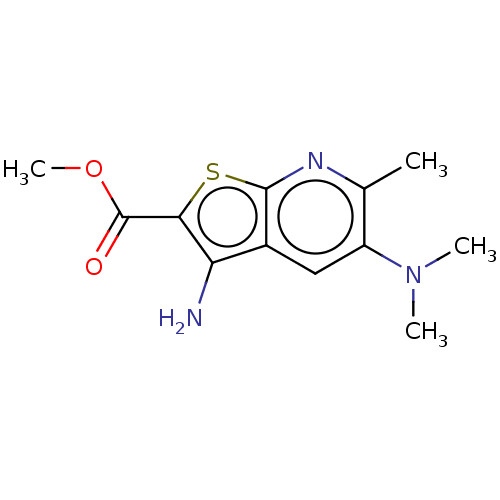
Affinity DataIC50: 150nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
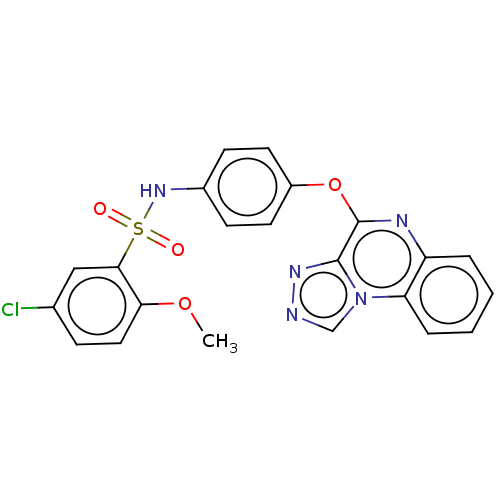
Affinity DataIC50: 160nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 200nMAssay Description:Inhibition of UT-B in Sprague-Dawley rat erythrocyte incubated for 6 min by erythrocyte osmotic lysis based high-throughput screening assayMore data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 220nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 270nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 280nMAssay Description:Inhibition of UT-B in Sprague-Dawley rat erythrocyte incubated for 6 min by erythrocyte osmotic lysis based high-throughput screening assayMore data for this Ligand-Target Pair
Affinity DataIC50: 280nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 430nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 470nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 530nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 540nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 600nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 610nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 640nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 660nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 710nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 750nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 800nMAssay Description:Inhibition of rat UT-B incubated for 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 800nMAssay Description:Inhibition of UT-B in rat erythrocyte by stopped-flow light scattering assayMore data for this Ligand-Target Pair
Affinity DataIC50: 810nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 870nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 900nMAssay Description:Inhibition of rat UT-B expressed in MDCK cells assessed as inhibition of urea transport incubated for 30 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 910nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.04E+3nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.06E+3nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.23E+3nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.41E+3nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 1.63E+3nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.66E+3nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Inhibition of rat UT-B incubated for 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.54E+3nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.77E+3nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Erythrocyte suspension (100 µL) was added to each well of a 96-well microplate to which test compounds were added. After 15 min incubation, 20 &...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of rat UT-B incubated for 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.17E+3nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair
Affinity DataIC50: 3.29E+3nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.36E+3nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.47E+3nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.51E+3nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.75E+3nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:Inhibition of rat UT-B incubated for 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 4.80E+3nMAssay Description:Erythrocyte suspension (100 µL) was added to each well of a 96-well microplate to which test compounds were added. After 15 min incubation, 20 &...More data for this Ligand-Target Pair
Affinity DataIC50: 5.46E+3nMAssay Description:Inhibition of urea transporter B in Sprague-Dawley rat erythrocytes incubated for 6 mins by spectrophotometric analysis based erythrocyte lysis assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMAssay Description:Inhibition of rat UT-B incubated for 15 minsMore data for this Ligand-Target Pair
Affinity DataIC50: 6.05E+3nMAssay Description:Inhibition of rat UT-B expressed in MDCK cellsMore data for this Ligand-Target Pair