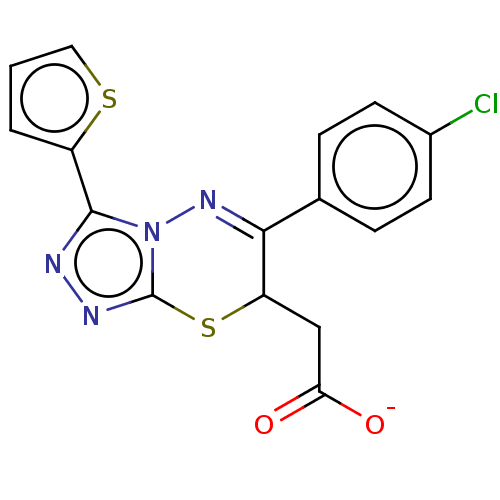
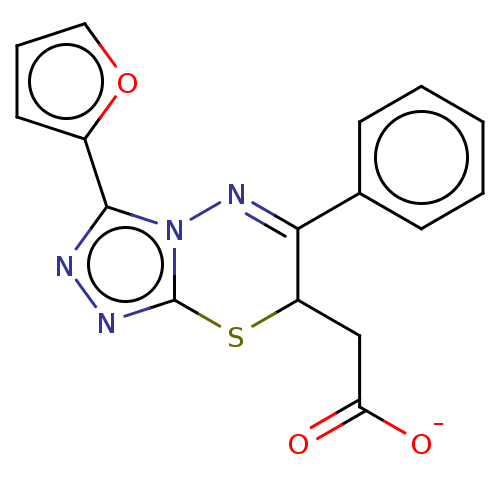
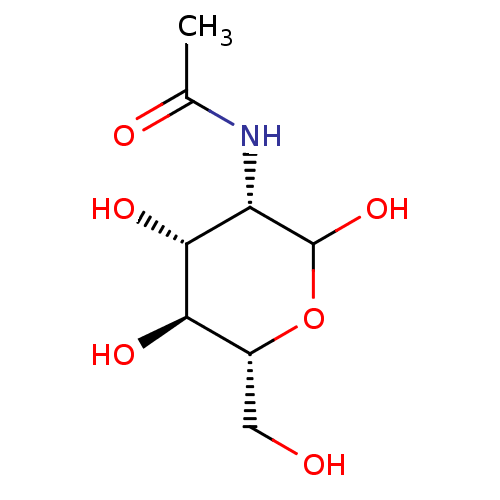
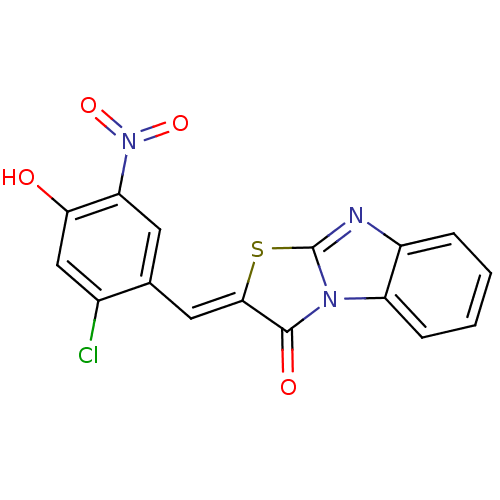
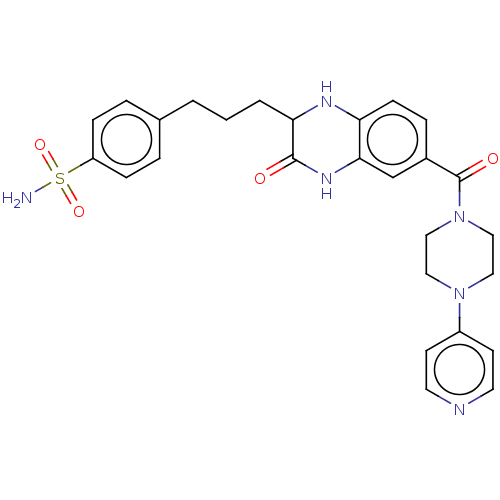
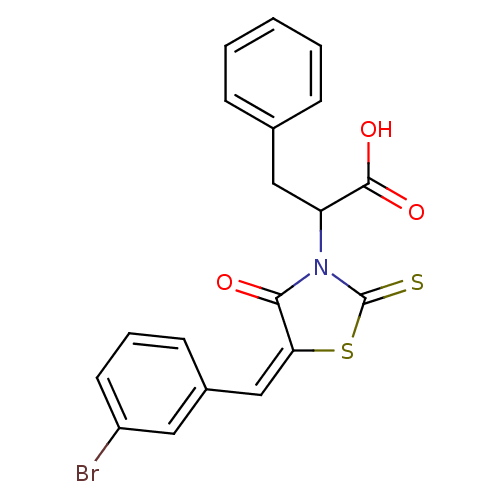
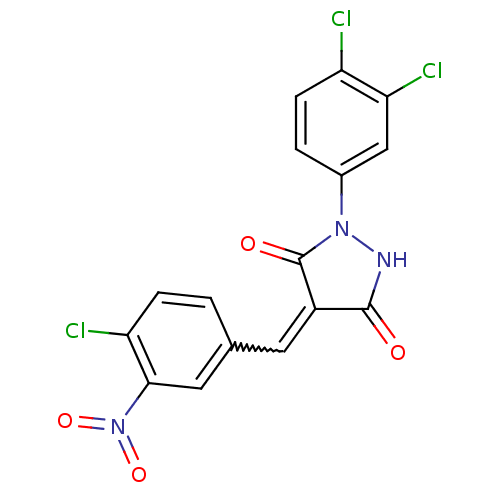
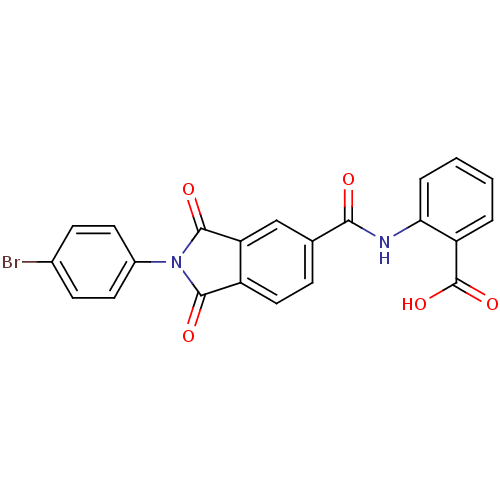
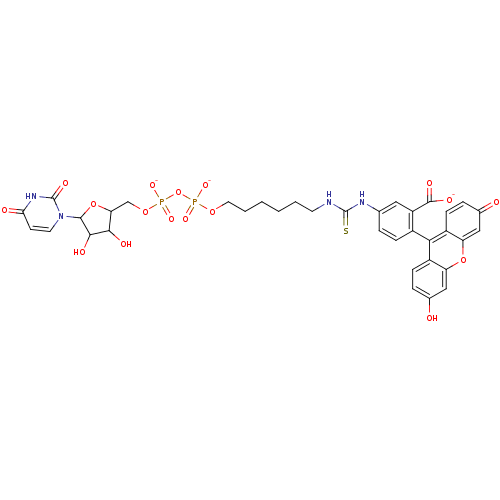
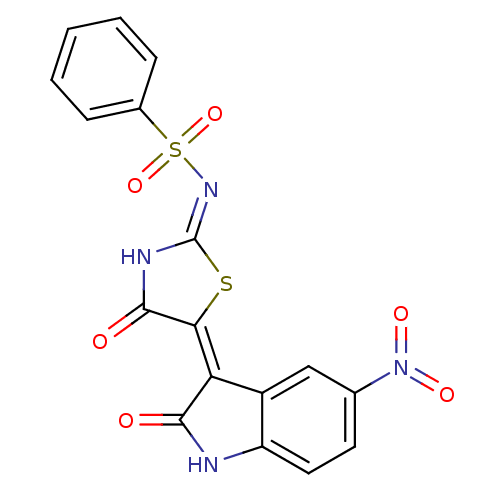
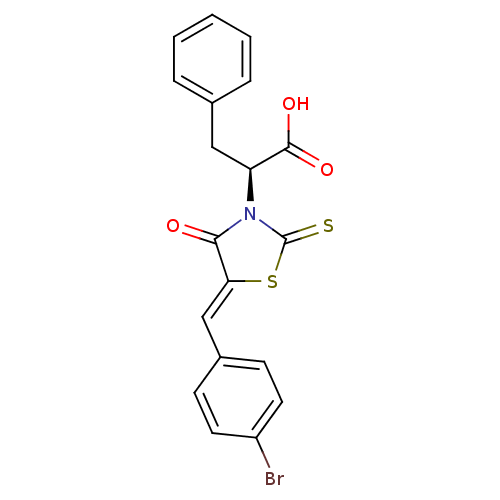
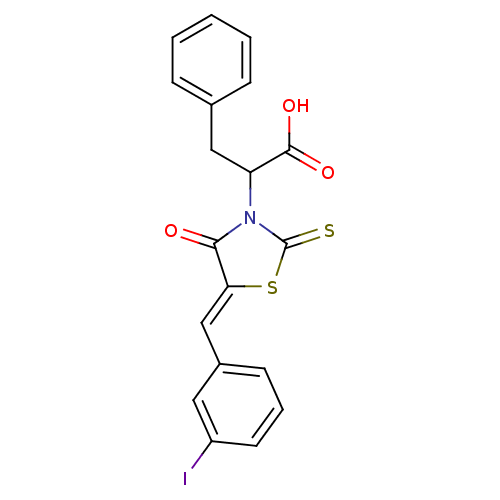
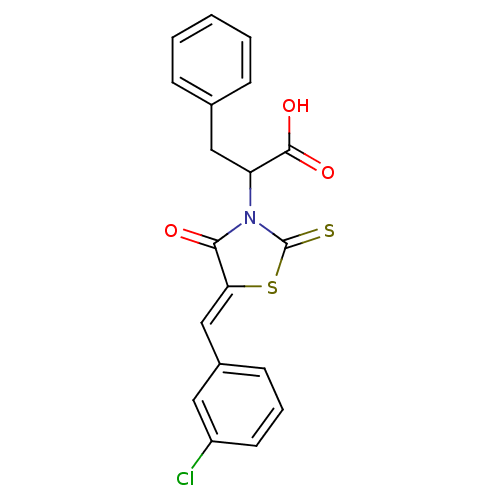
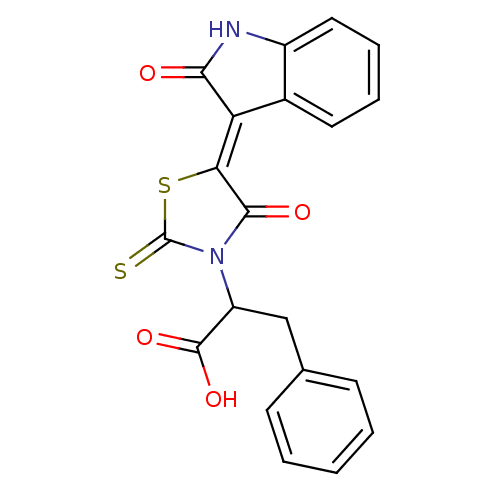
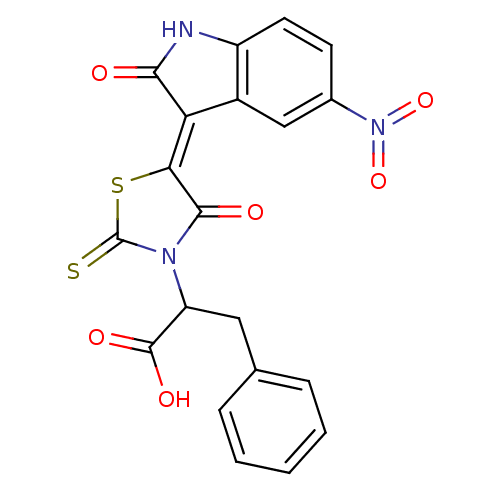
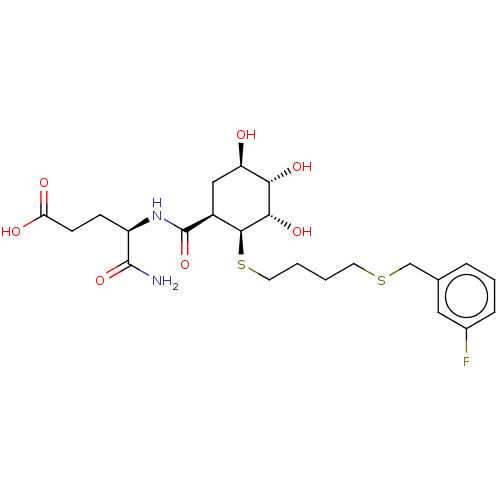
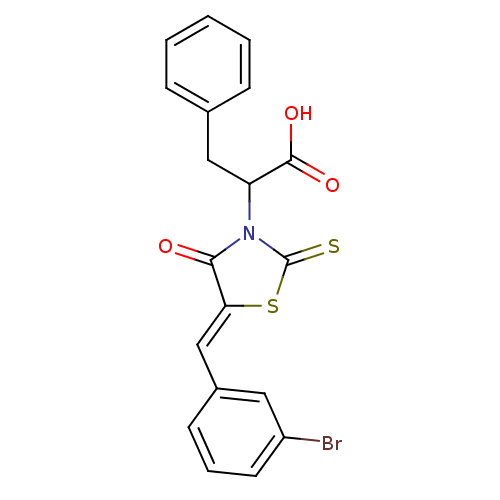
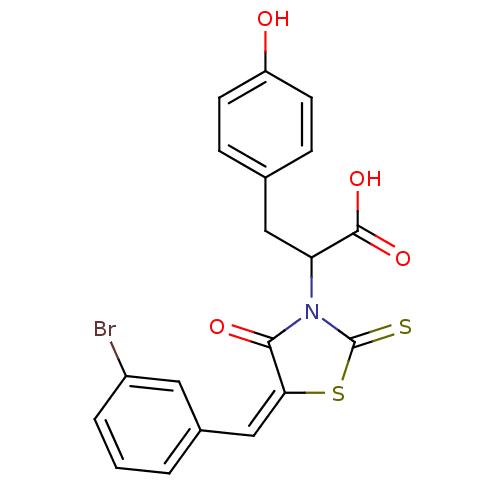
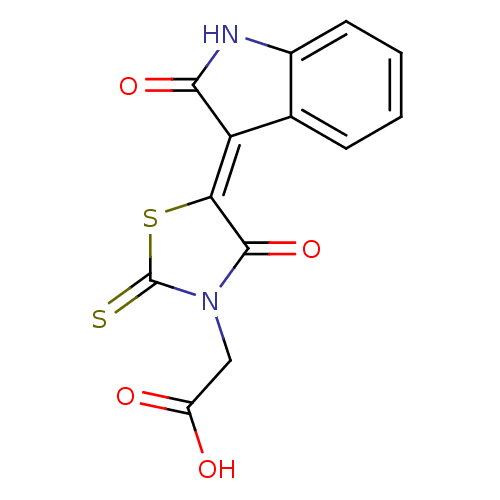
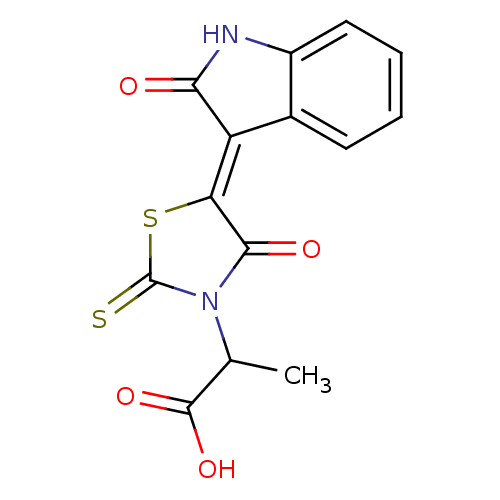
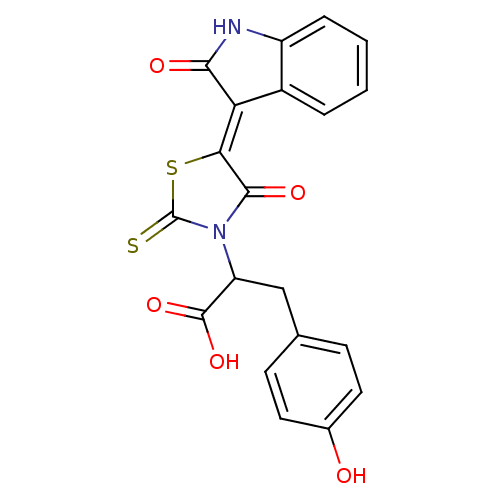
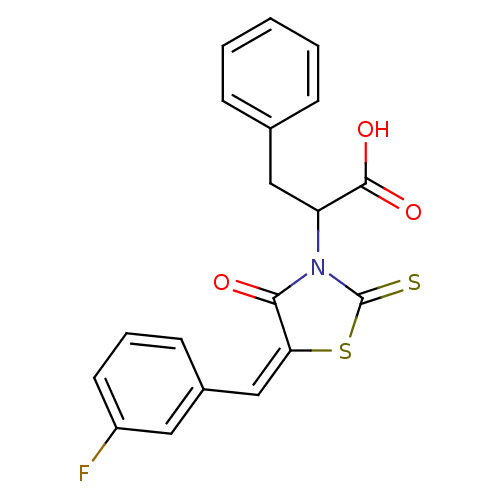
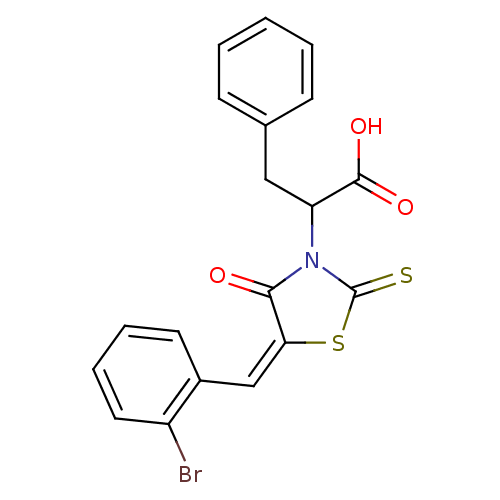
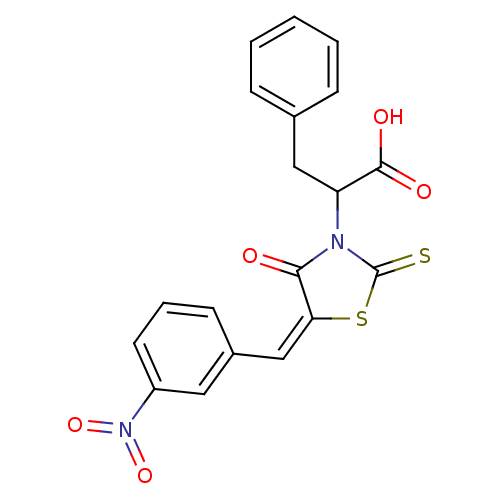
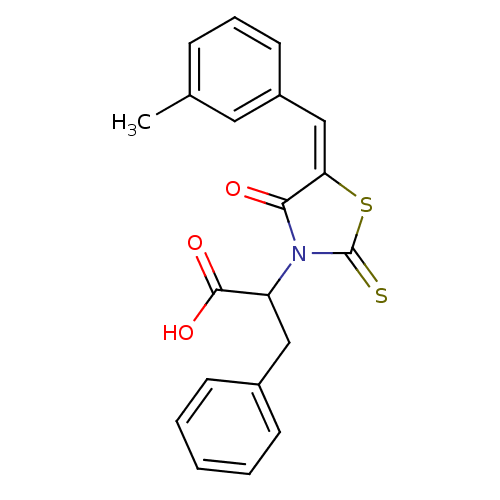
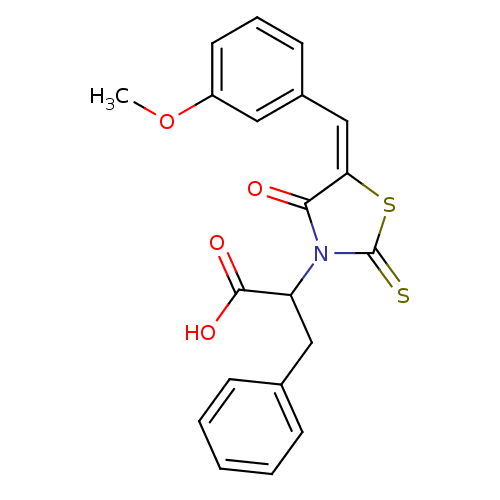
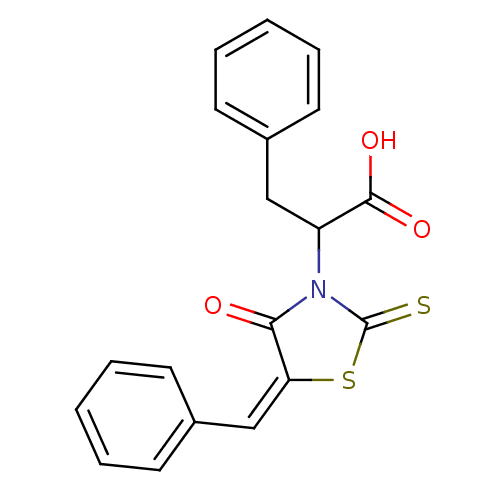
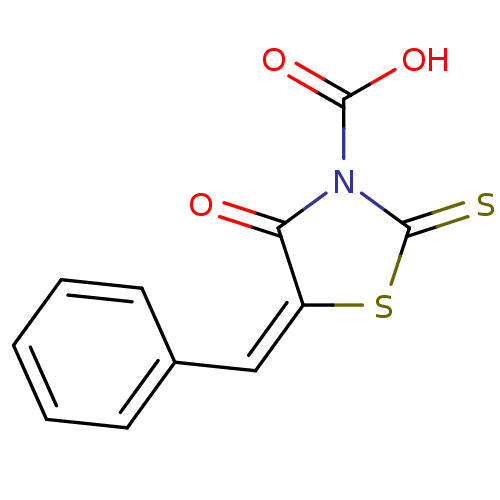
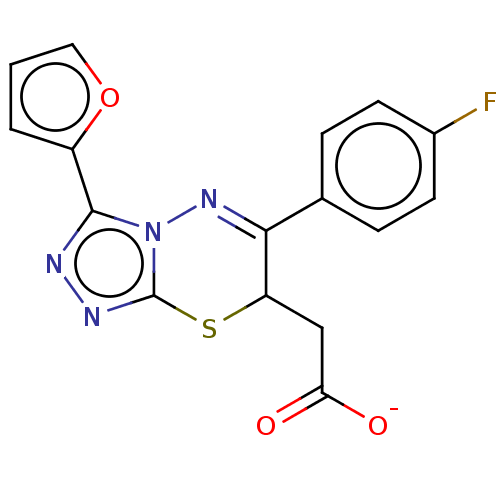
TargetProbable UDP-galactopyranose mutase(Klebsiella pneumoniae (Enterobacteria))
University of Wisconsin-Madison
University of Wisconsin-Madison
Affinity DataKi: 8.00E+3nM ΔG°: -28.8kJ/moleT: 2°CAssay Description:Reactions were performed in duplicate at 22�C in 50 mM NaH2PO4 buffer (pH 7) with 20 nM enzyme and 20 mM fresh sodium dithionite. Small molecules wer...More data for this Ligand-Target Pair
Affinity DataKi: 2.50E+4nM ΔG°: -26.0kJ/moleT: 2°CAssay Description:Reactions were performed in duplicate at 22�C in 50 mM NaH2PO4 buffer (pH 7) with 20 nM enzyme and 20 mM fresh sodium dithionite. Small molecules wer...More data for this Ligand-Target Pair
Affinity DataKi: 3.10E+4nM ΔG°: -25.5kJ/moleT: 2°CAssay Description:Reactions were performed in duplicate at 22�C in 50 mM NaH2PO4 buffer (pH 7) with 20 nM enzyme and 20 mM fresh sodium dithionite. Small molecules wer...More data for this Ligand-Target Pair
Affinity DataKi: 7.70E+4nM ΔG°: -23.2kJ/moleT: 2°CAssay Description:Reactions were performed in duplicate at 22�C in 50 mM NaH2PO4 buffer (pH 7) with 20 nM enzyme and 20 mM fresh sodium dithionite. Small molecules wer...More data for this Ligand-Target Pair
TargetProbable UDP-galactopyranose mutase(Klebsiella pneumoniae (Enterobacteria))
University of Wisconsin-Madison
University of Wisconsin-Madison
Affinity DataKi: 7.80E+4nM ΔG°: -23.2kJ/moleT: 2°CAssay Description:Reactions were performed in duplicate at 22�C in 50 mM NaH2PO4 buffer (pH 7) with 20 nM enzyme and 20 mM fresh sodium dithionite. Small molecules wer...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+7nMAssay Description:Inhibition of DC-SIGN (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 28nM Kd: 5.30E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:Inhibition of DC-SIGN (unknown origin) after 1 hr by fluorescence analysisMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nM Kd: 9.40E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nM Kd: 9.80E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70E+3nM Kd: 160nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+4nM Kd: 4.00E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+4nM Kd: 9.30E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+4nM Kd: 6.00E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 4.10E+4nM Kd: 4.90E+3nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+4nM Kd: 1.60E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 6.50E+4nM Kd: 1.20E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 9.30E+4nM Kd: 1.07E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: >1.00E+5nM Kd: 1.00E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+6nMAssay Description:Inhibition of DC-SIGN (unknown origin)More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >1.12E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: 1.41E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+5nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+5nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+5nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+4nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: >3.30E+5nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
Affinity DataKd: 100nMAssay Description:To access the ability of compounds to inhibit the catalytic activity of UGM, an HPLC assay was employed. Inhibition constants were determined by mon...More data for this Ligand-Target Pair
TargetProbable UDP-galactopyranose mutase(Klebsiella pneumoniae (Enterobacteria))
University of Wisconsin-Madison
University of Wisconsin-Madison
Affinity DataKd: 4.10E+4nMT: 2°CAssay Description:Reactions were performed in duplicate at 22�C in 50 mM NaH2PO4 buffer (pH 7) with 20 nM enzyme and 20 mM fresh sodium dithionite. Small molecules wer...More data for this Ligand-Target Pair
TargetProbable UDP-galactopyranose mutase(Klebsiella pneumoniae (Enterobacteria))
University of Wisconsin-Madison
University of Wisconsin-Madison
Affinity DataKd: >1.00E+5nMT: 2°CAssay Description:Reactions were performed in duplicate at 22�C in 50 mM NaH2PO4 buffer (pH 7) with 20 nM enzyme and 20 mM fresh sodium dithionite. Small molecules wer...More data for this Ligand-Target Pair