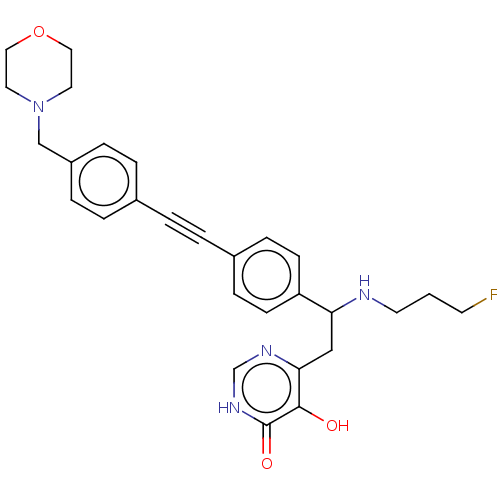
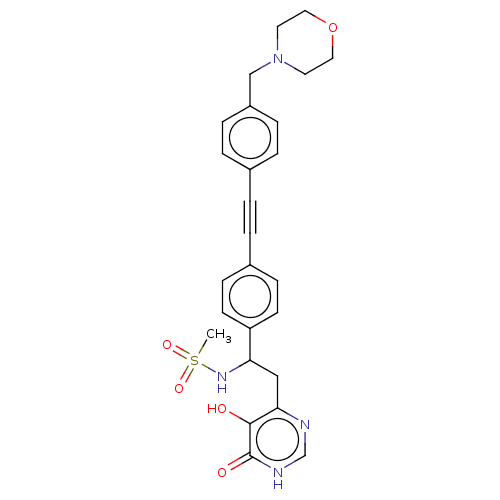
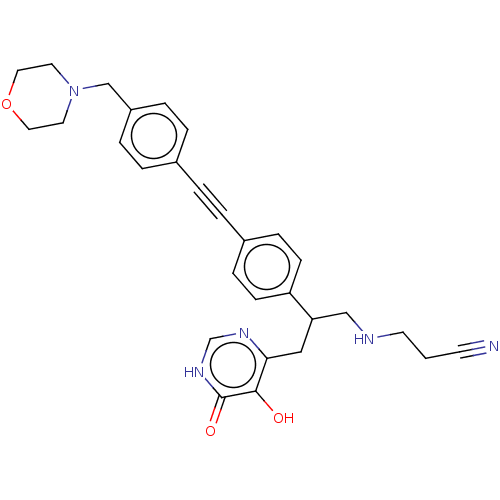
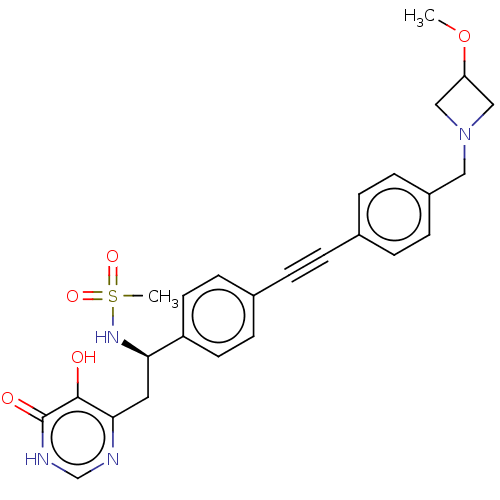
Report error Found 61 Enz. Inhib. hit(s) with all data for entry = 10745
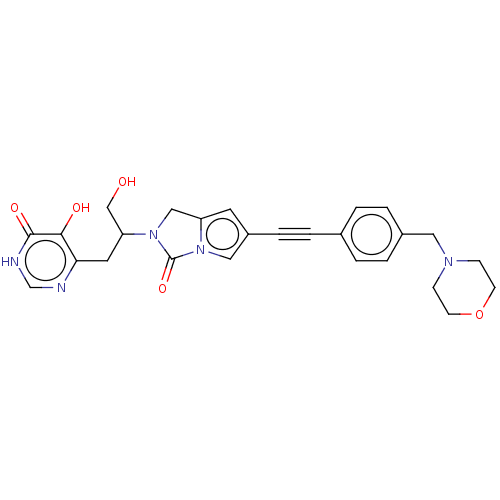
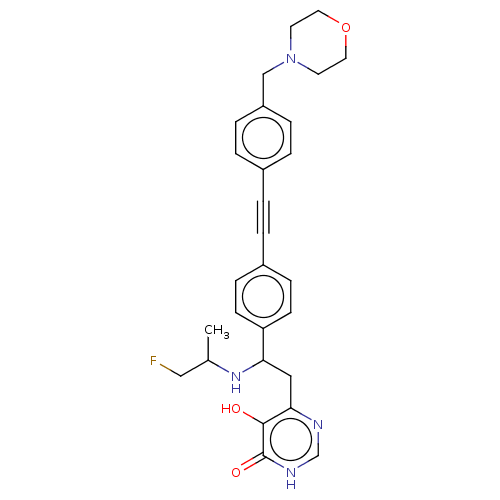
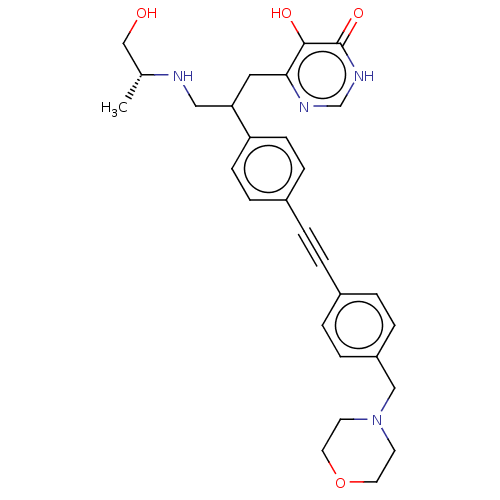
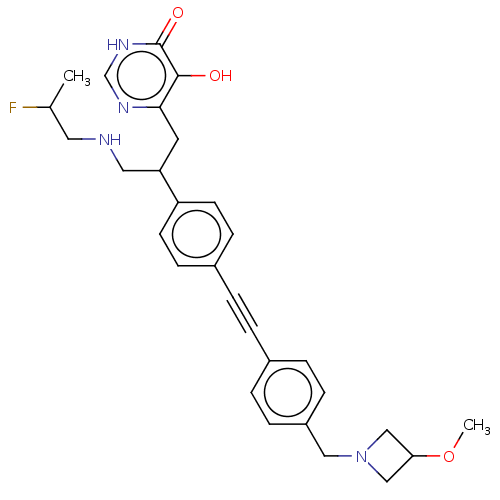
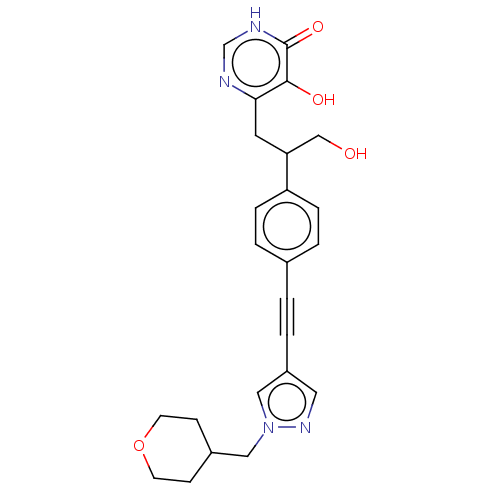
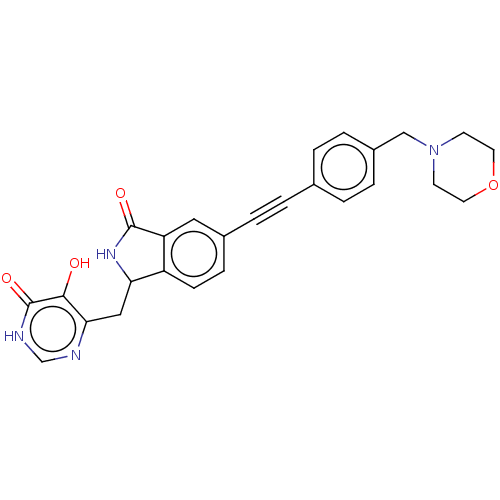
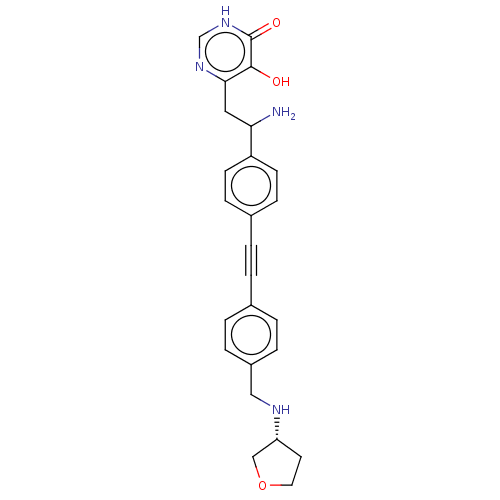
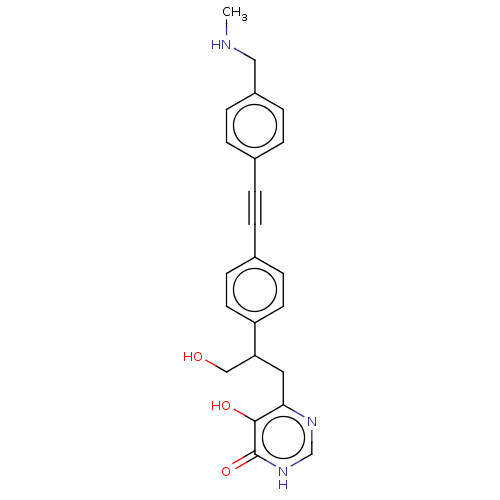
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
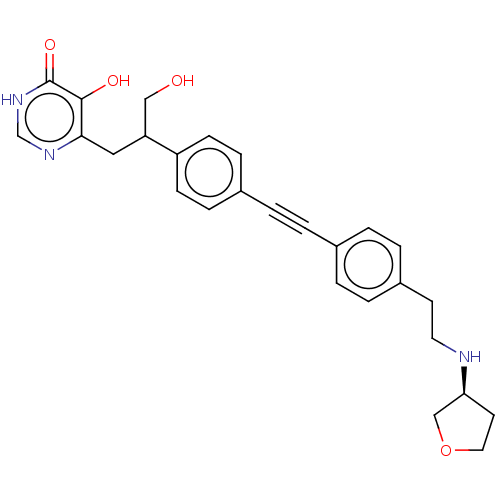
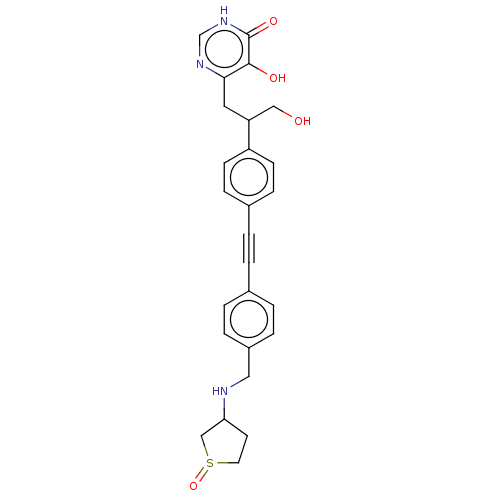
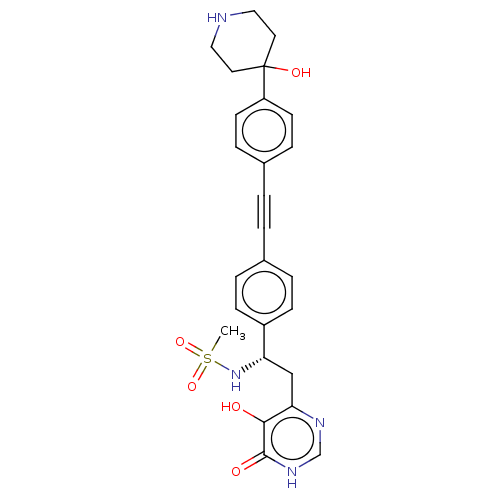
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
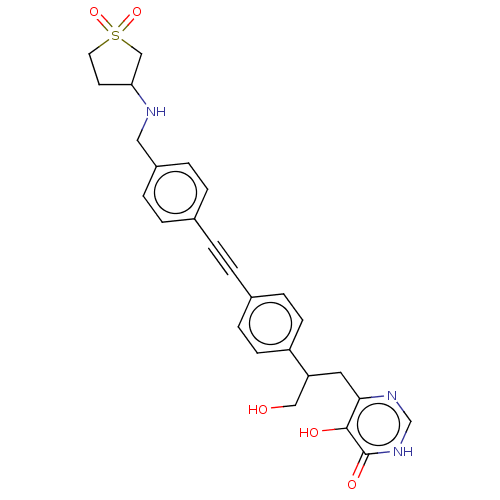
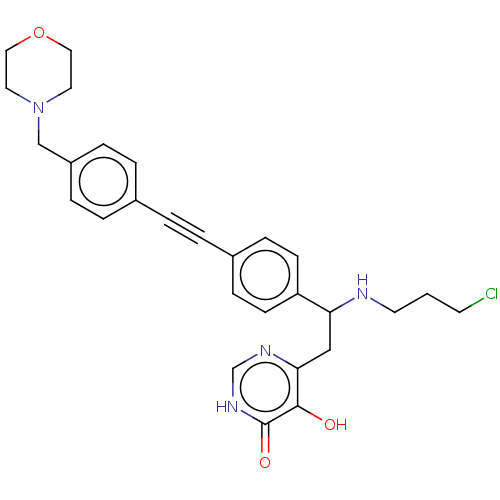
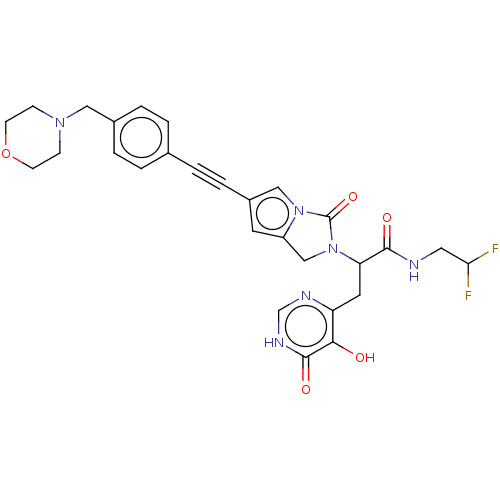
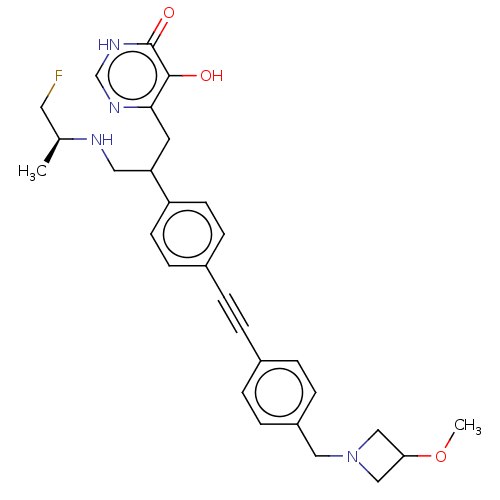
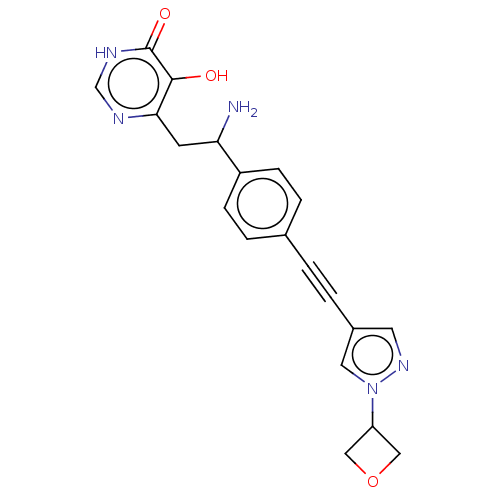
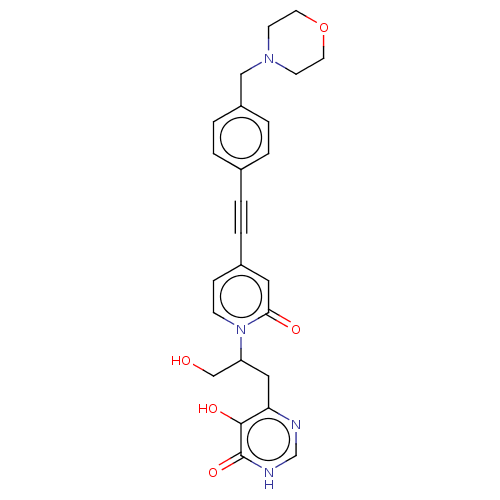
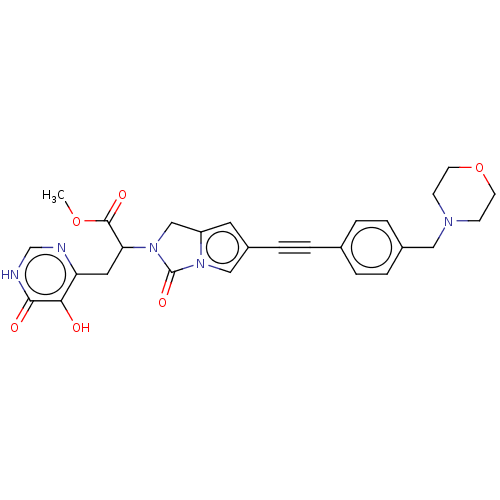
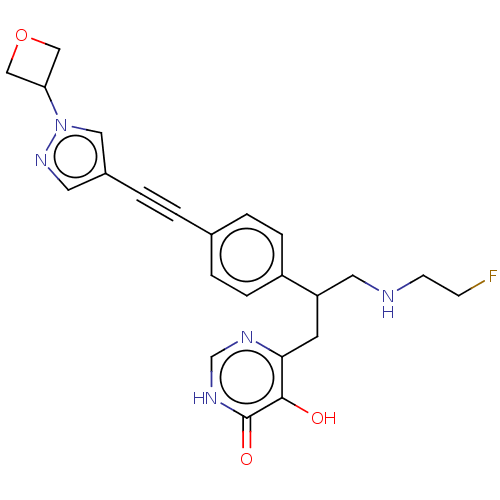
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
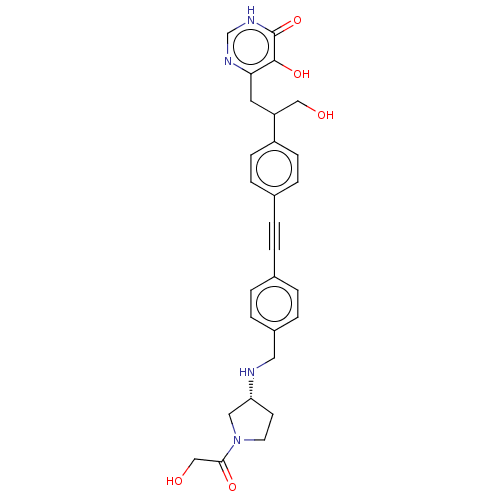
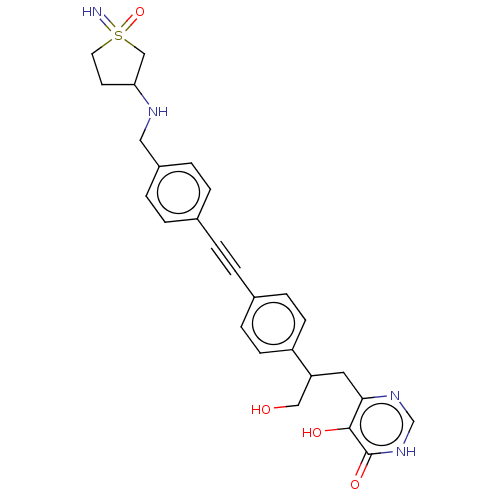
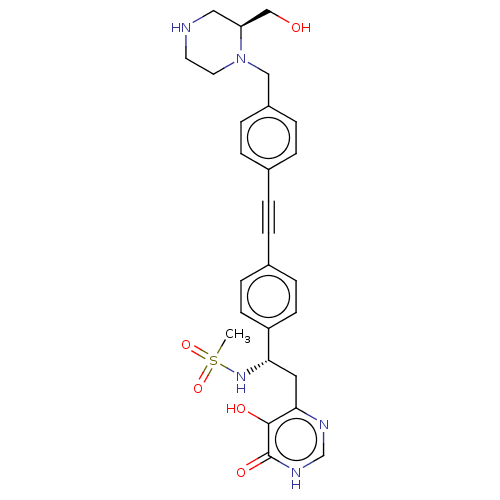
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
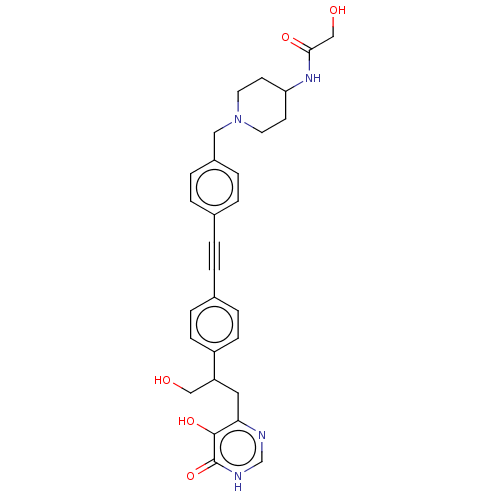
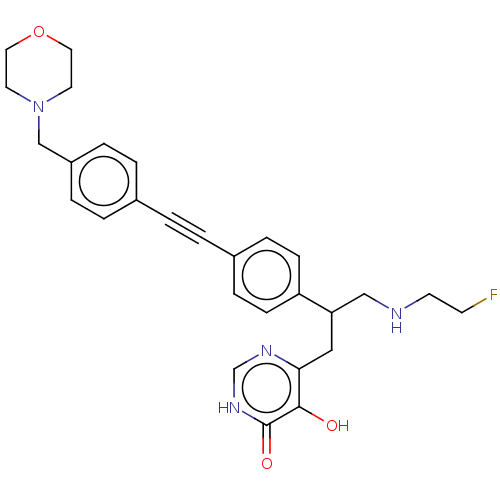
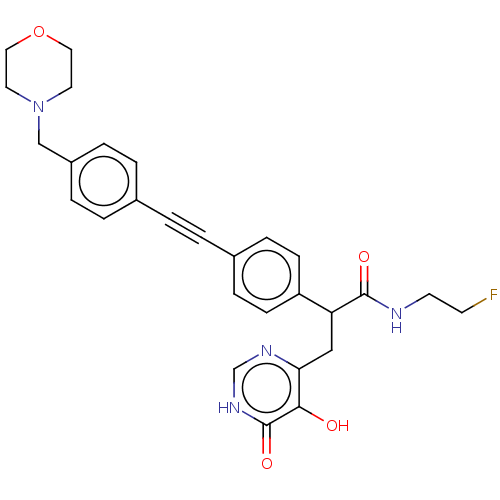
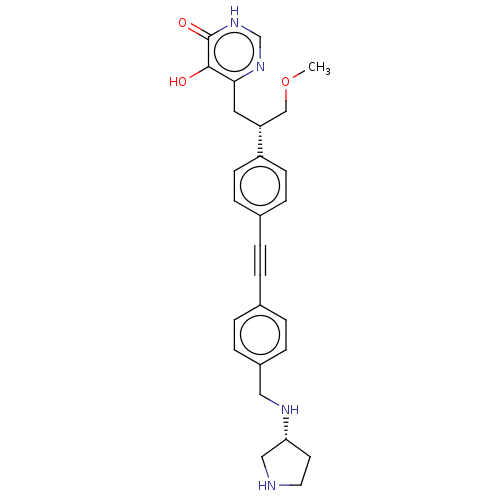
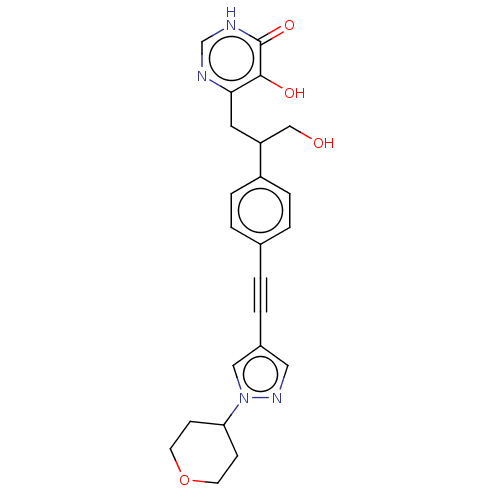
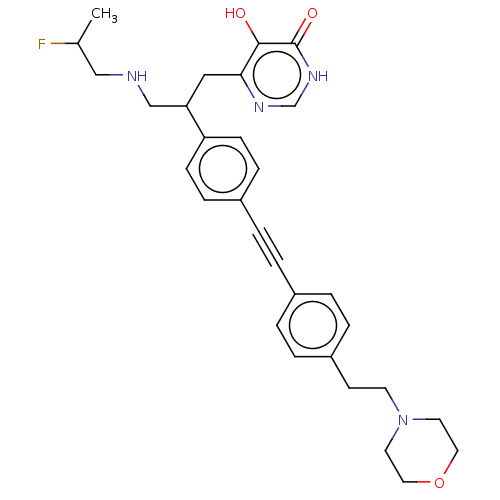
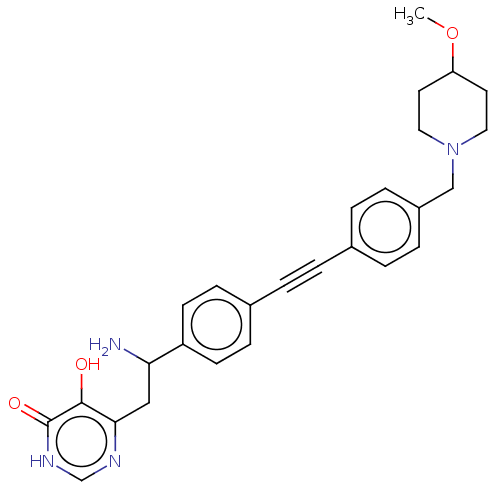
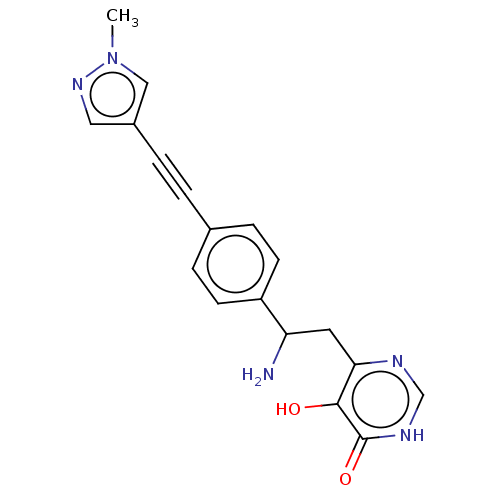
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
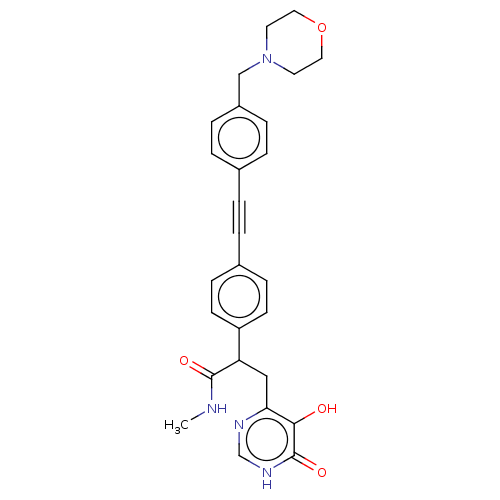
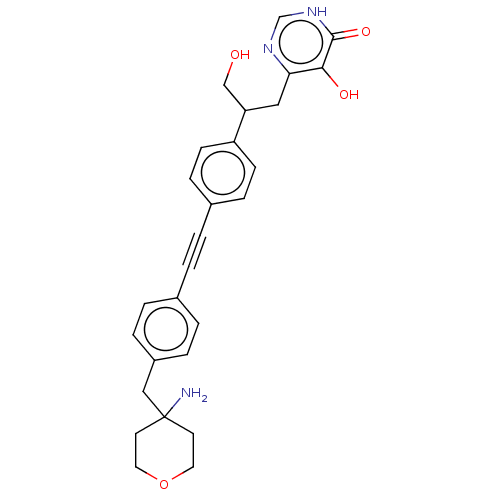
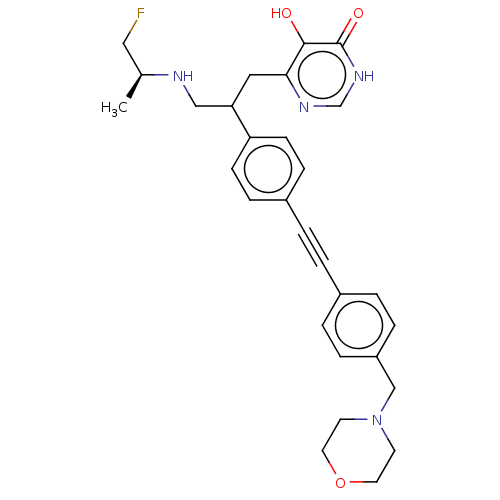
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
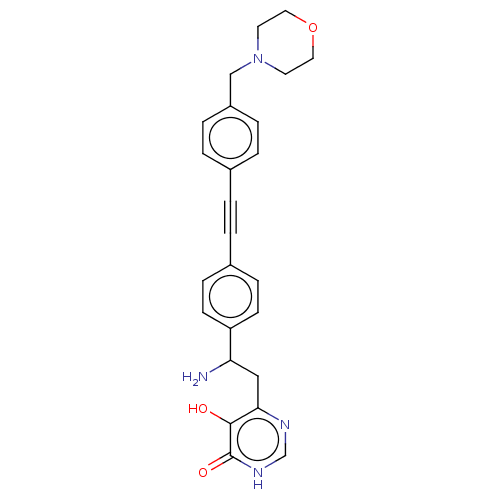
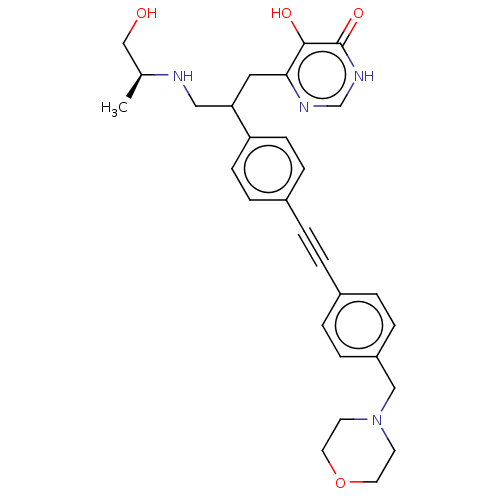
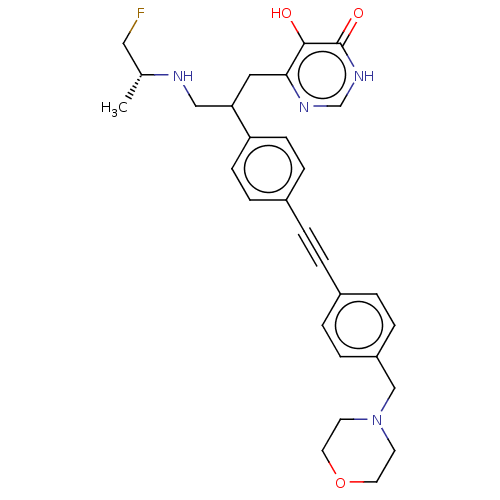
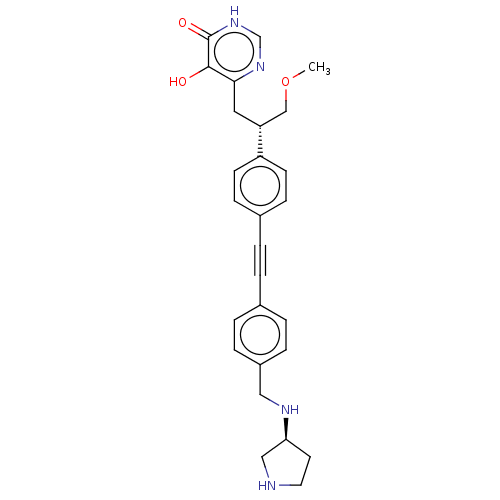
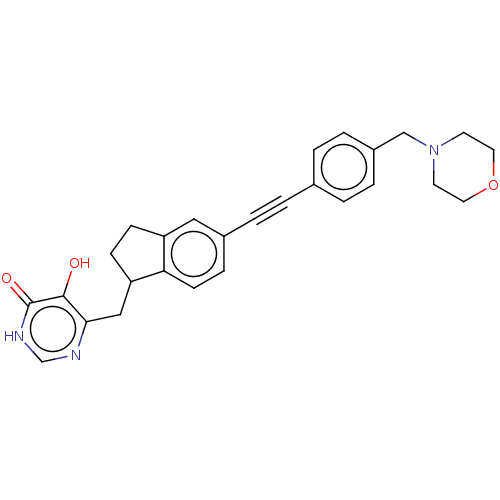
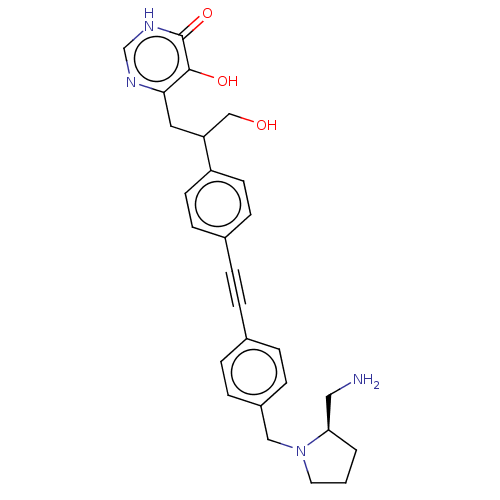
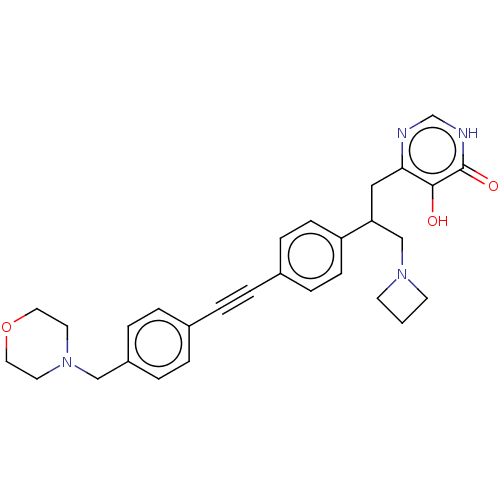
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
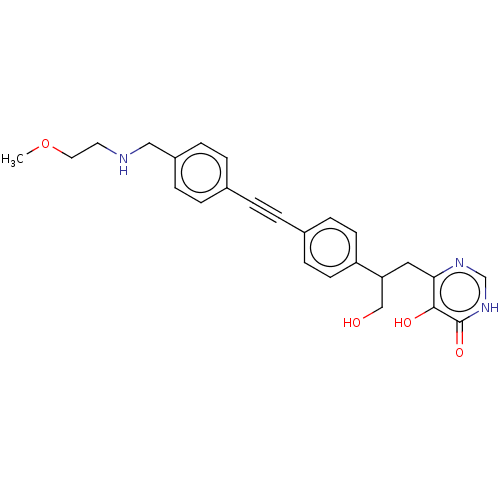
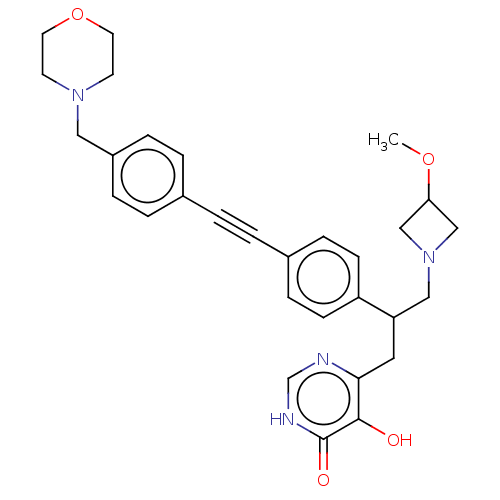
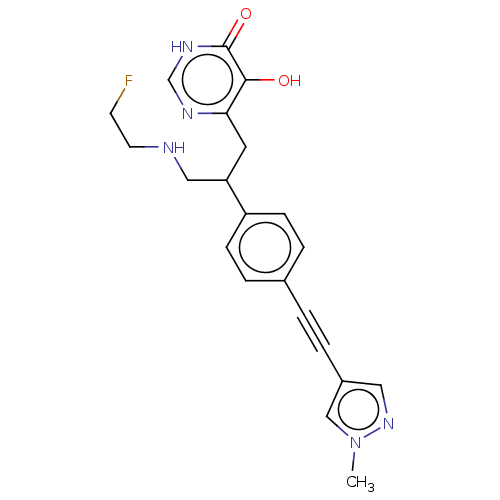
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 10nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 30nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 75nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 75nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Forge Therapeutics
US Patent
Forge Therapeutics
US Patent
Affinity DataIC50: 550nMAssay Description:IC50 values against E. coli LpxC were determined using a Rapid Fire MS assay as previously described J. Med. Chem. 2012, 55, 1662-1670.More data for this Ligand-Target Pair