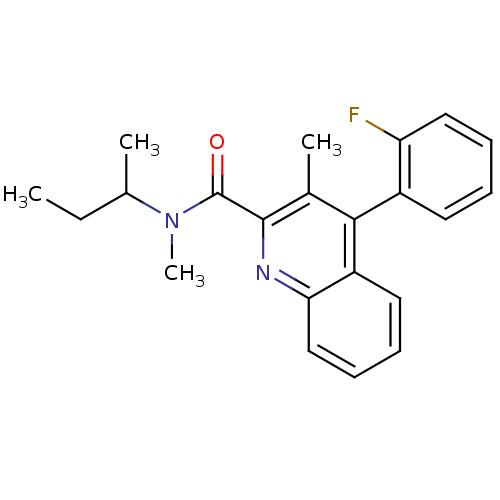
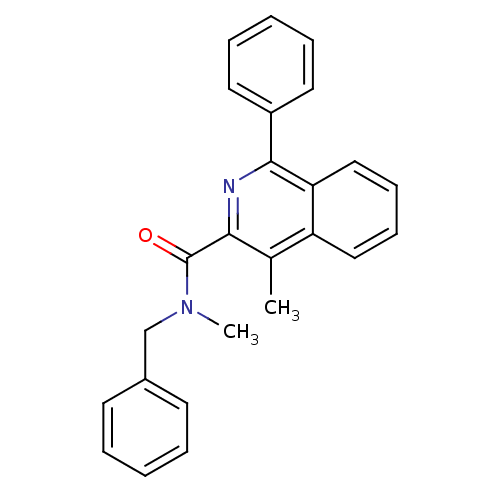
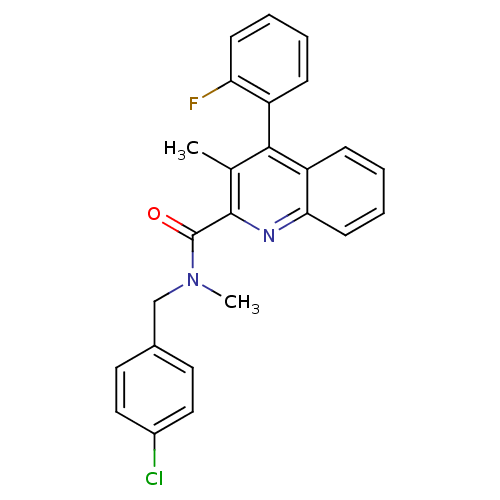
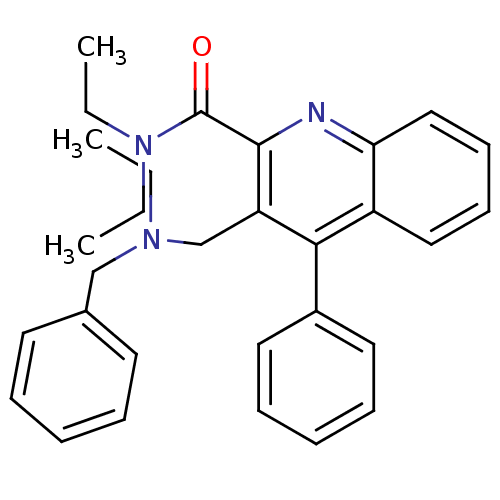
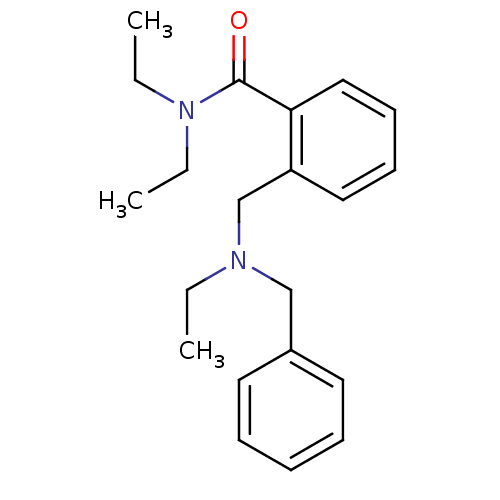
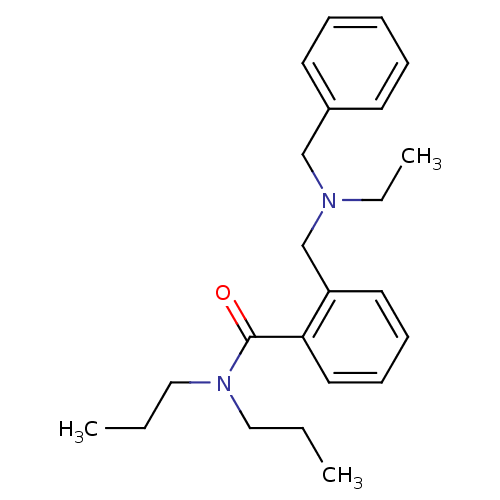
Report error Found 53 Enz. Inhib. hit(s) with all data for entry = 50037009
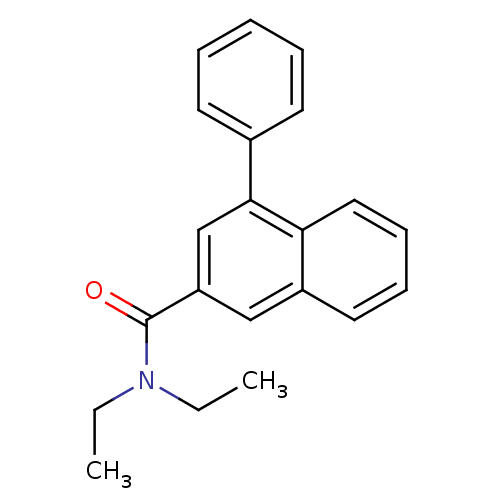
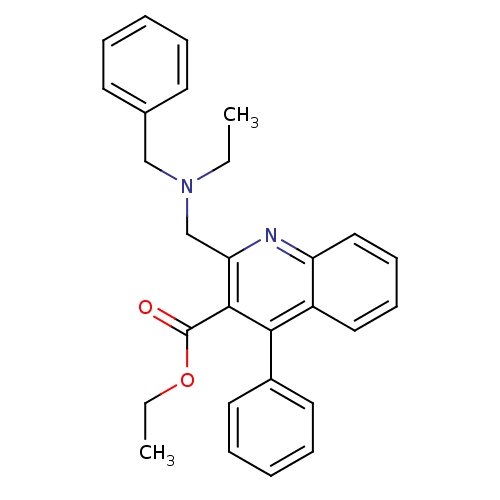
Affinity DataIC50: 2.10nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
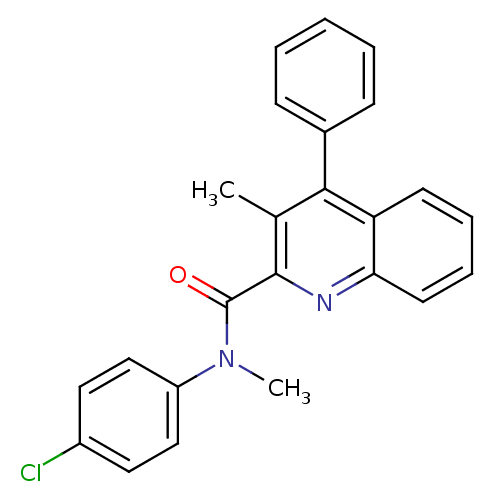
Affinity DataIC50: 2.10nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
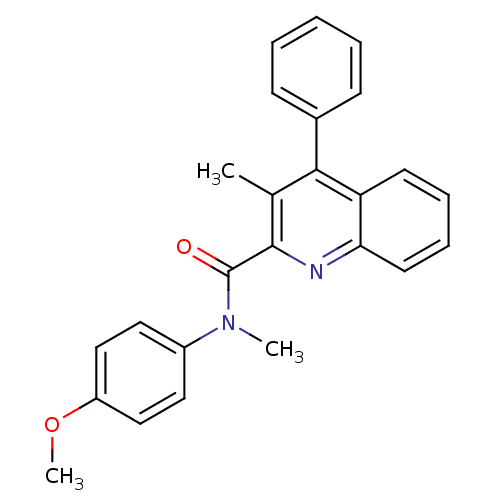
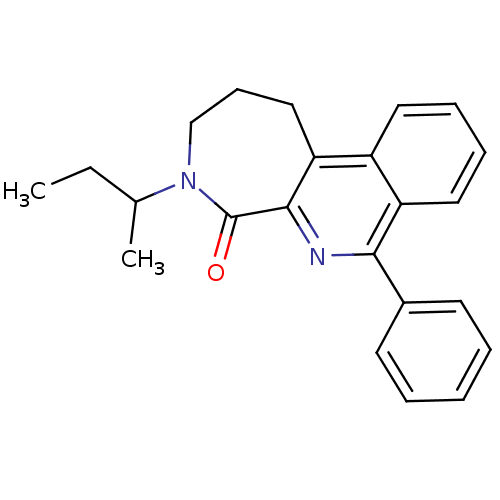
Affinity DataIC50: 2.20nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
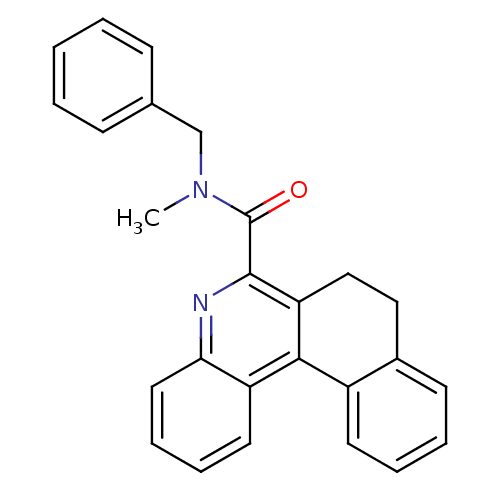
Affinity DataIC50: 2.20nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
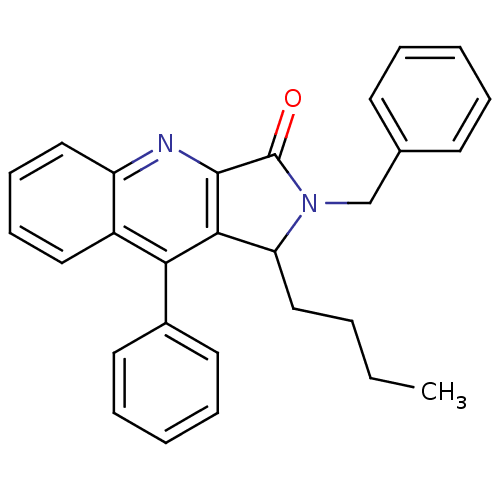
Affinity DataIC50: 2.90nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
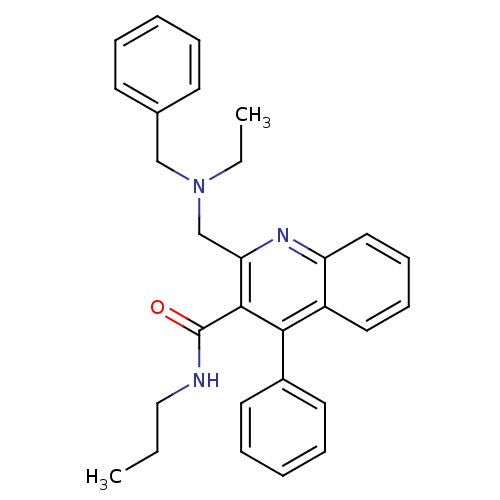
Affinity DataIC50: 3.10nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 3.40nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 4.10nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 6.40nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 8.80nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 8.90nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 9.80nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 45nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 170nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 210nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 229nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 270nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 373nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 410nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 456nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 480nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 490nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 540nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 550nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 620nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 700nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 780nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 810nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 872nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 1.19E+3nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20E+3nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 1.32E+3nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 2.48E+3nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+3nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 3.40E+3nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 3.70E+3nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 8.21E+3nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 8.87E+3nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Test in vitro, for potential ability to displace [3H]1 from PBR in rat brain cortexMore data for this Ligand-Target Pair
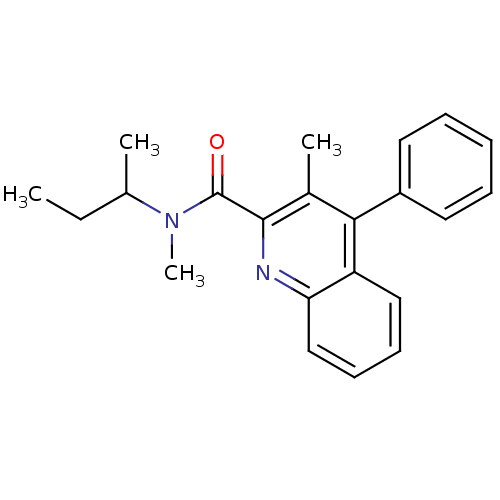
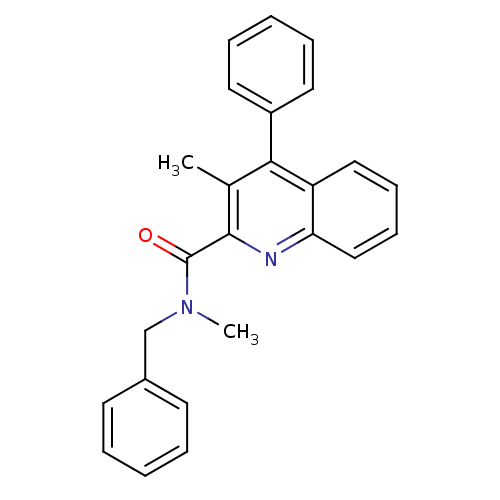
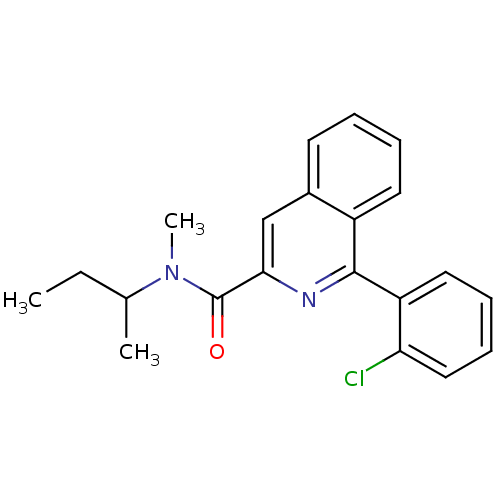
 3D Structure (crystal)
3D Structure (crystal)