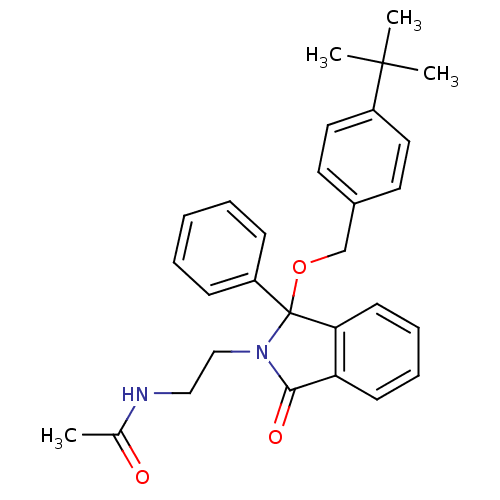
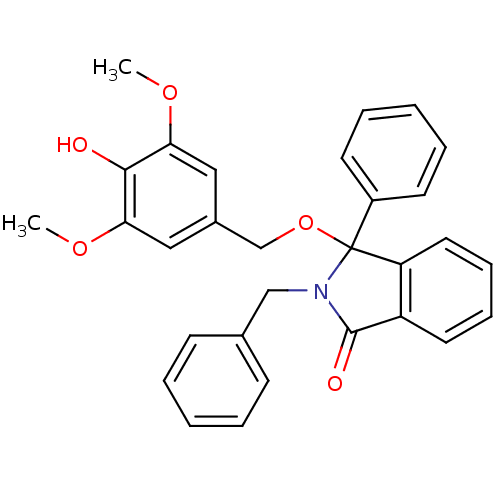
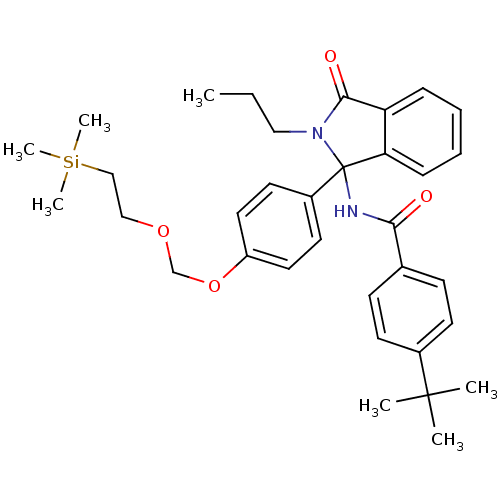
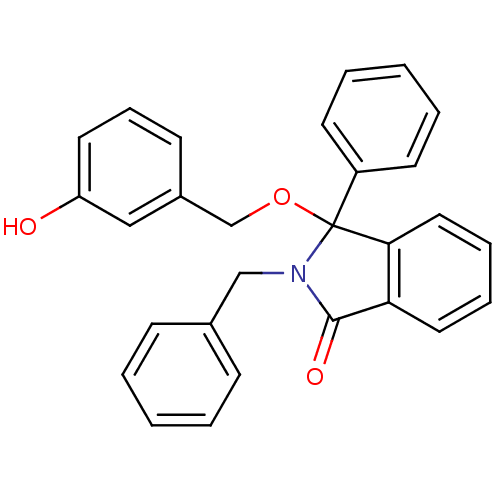
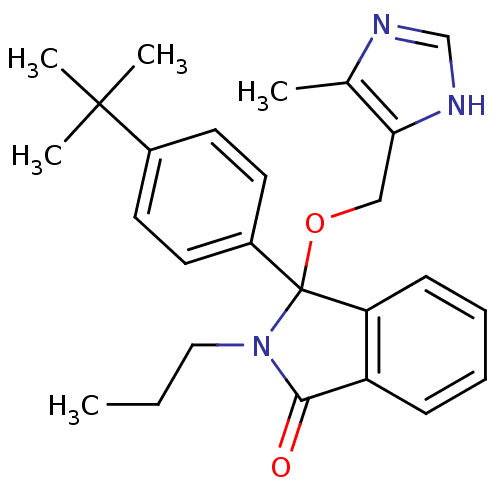
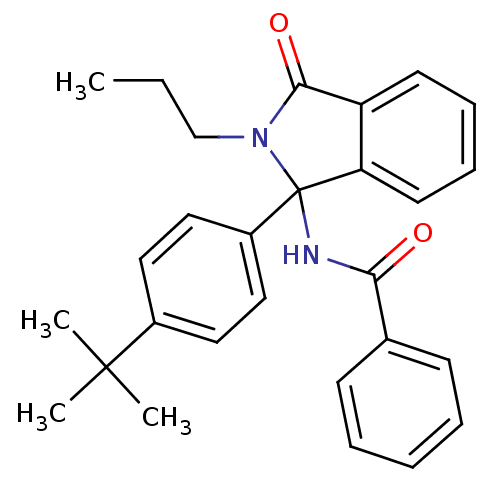
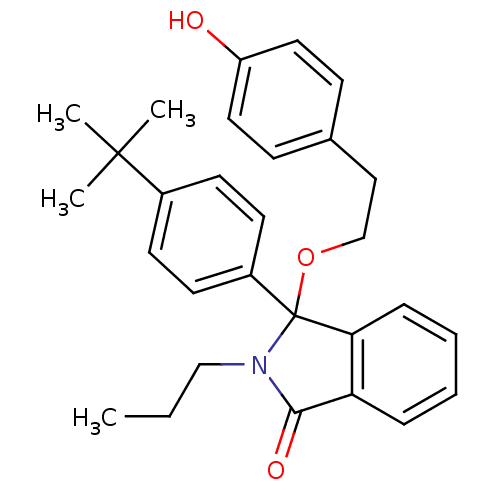
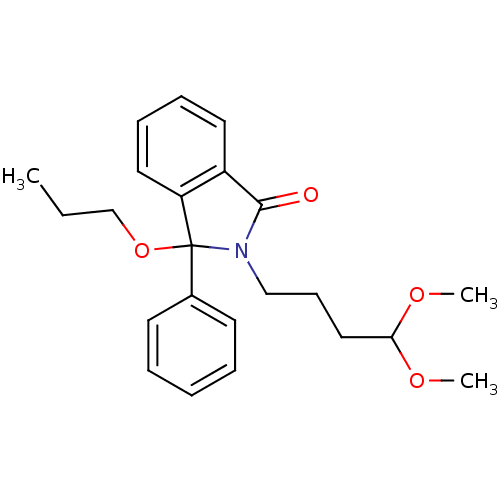
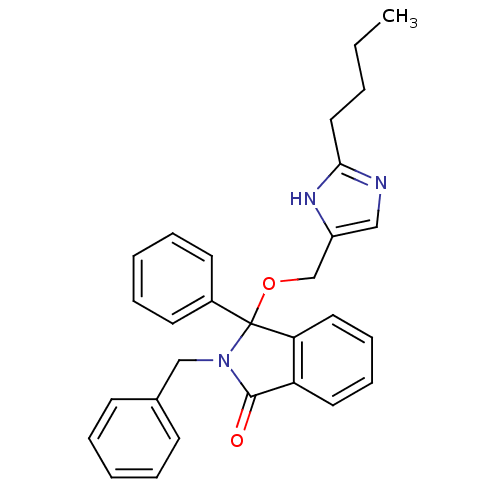
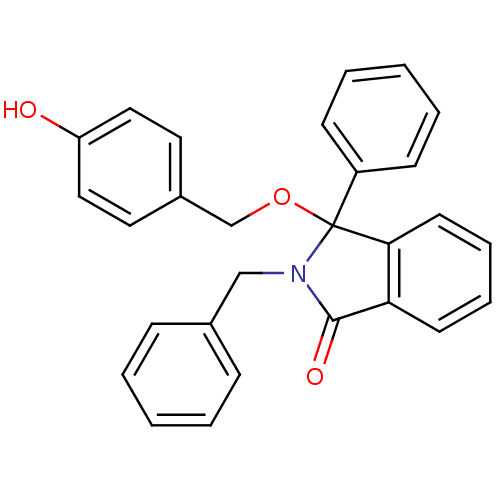
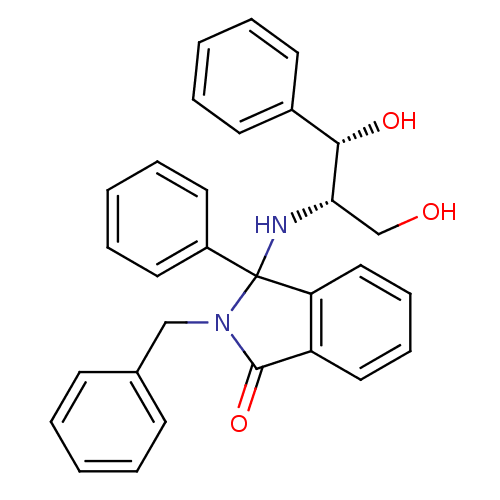
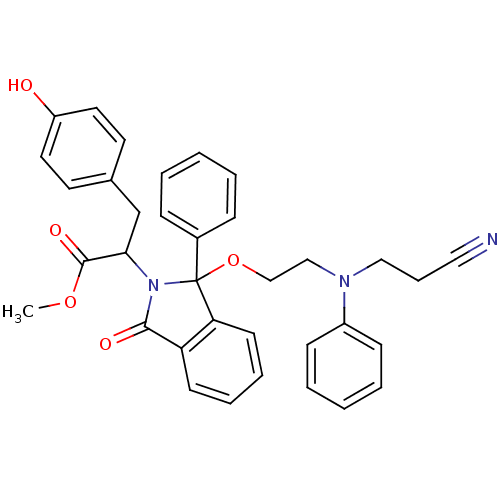
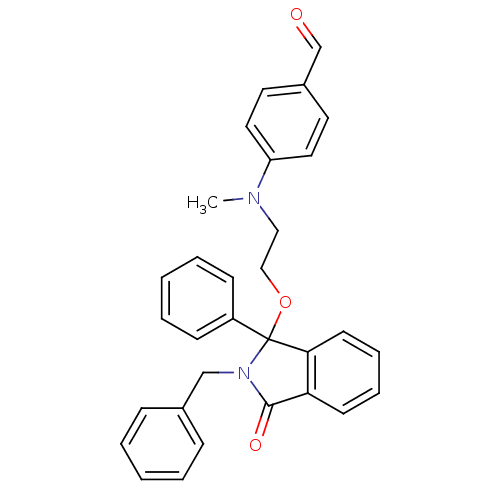
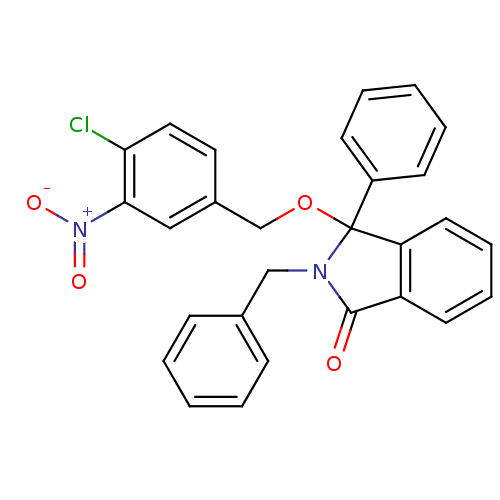
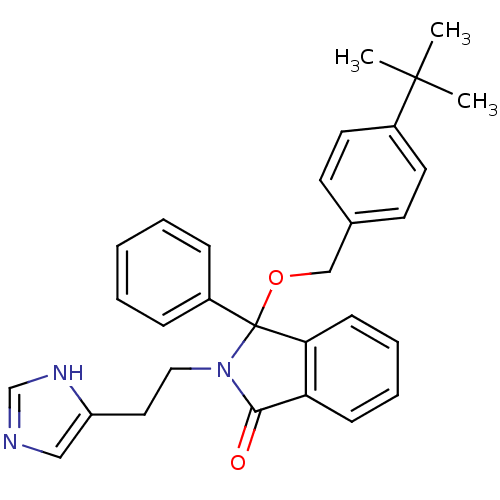
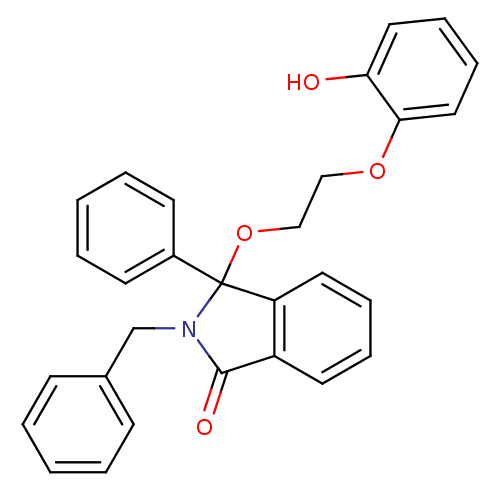
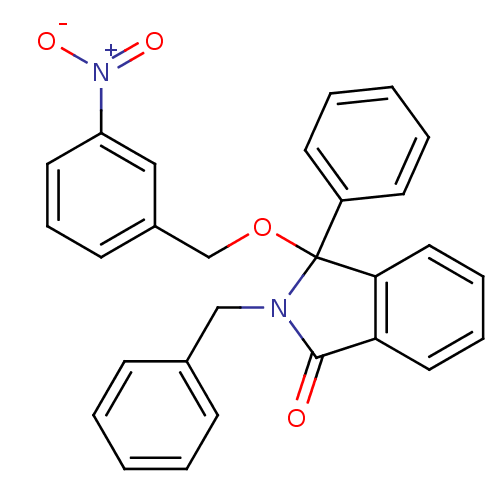
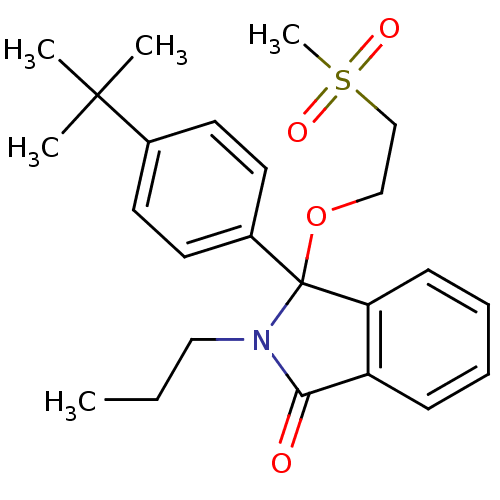
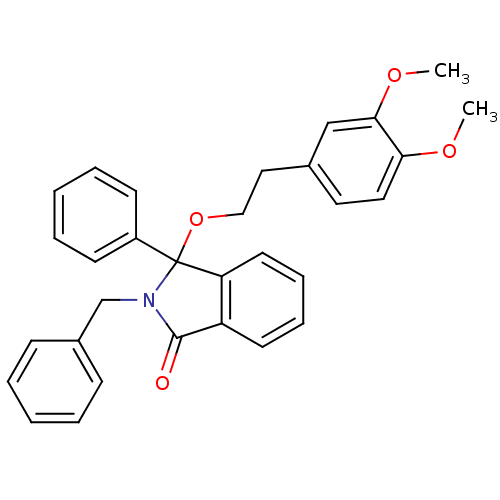
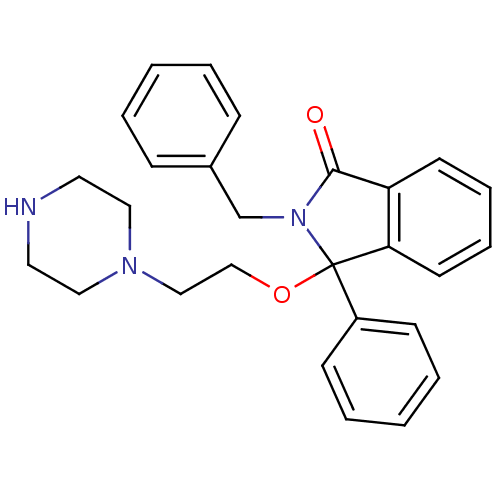
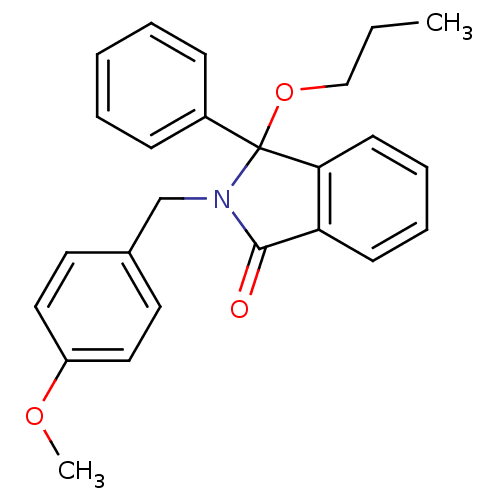
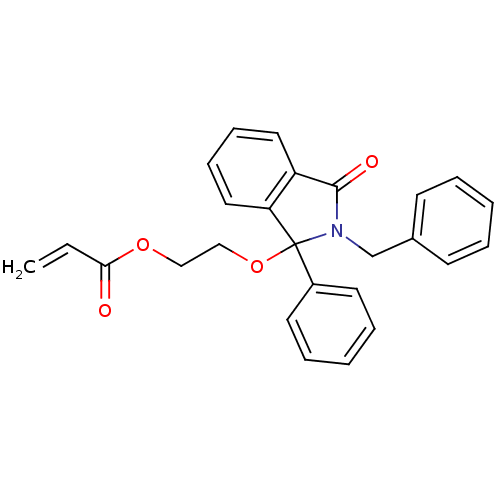
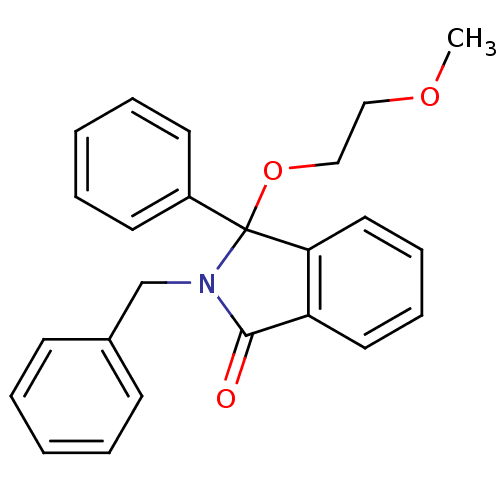
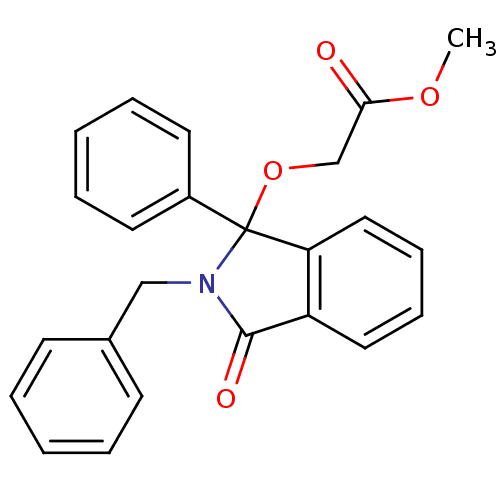
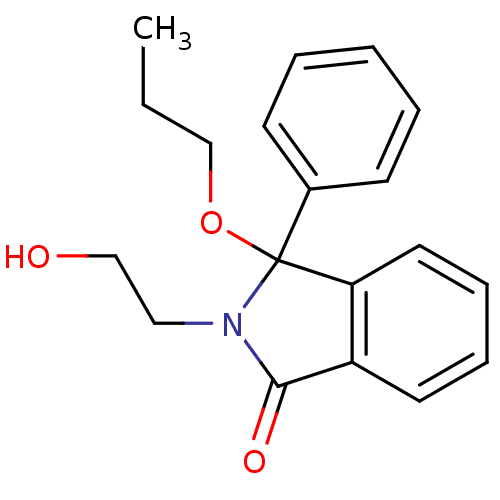
Report error Found 45 Enz. Inhib. hit(s) with all data for entry = 3312
Affinity DataIC50: 1.44E+4nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.79E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 2.70E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 6.43E+4nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 6.60E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 6.97E+4nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+4nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 7.01E+4nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 7.02E+4nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 7.43E+4nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 7.80E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 7.90E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 8.50E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 8.80E+4nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 9.00E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 9.20E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 9.60E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 9.70E+4nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.03E+5nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.03E+5nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.16E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.81E+5nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 1.91E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 2.06E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 2.14E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 2.21E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 2.32E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 2.43E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 2.75E+5nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 2.81E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 2.84E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+5nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 3.11E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 3.15E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 3.26E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 3.45E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 3.80E+5nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 3.93E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 4.13E+5nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 4.39E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 4.90E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMpH: 7.4 T: 2°CAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+5nMAssay Description:For initial testing, the compounds and controls were plated out in triplicate into clear 96-well plates (Nunc) in 10 ul aliquots to give final concen...More data for this Ligand-Target Pair